---
title: "Staggered Difference-in-Differences"
subtitle: "Modern Methods for Heterogeneous Treatment Timing"
---
```{r}
#| label: setup
#| include: false
knitr::opts_chunk$set(
echo = TRUE, warning = FALSE, message = FALSE,
fig.width = 10, fig.height = 6, fig.align = "center"
)
library(dplyr)
library(tidyr)
library(ggplot2)
library(fixest)
theme_set(theme_minimal(base_size = 13))
set.seed(42)
```
# The DiD Revolution {.unnumbered}
Difference-in-differences (DiD) is the workhorse of policy evaluation. But recent research has revealed that the standard two-way fixed effects (TWFE) estimator can produce severely biased estimates when treatment timing varies across units [@goodman2021; @callaway2021; @roth2023].
This module covers the "credibility revolution" in DiD—understanding why TWFE fails with staggered adoption and how modern estimators fix the problem.
## The Classic Setup
In the canonical 2×2 DiD, we have:
- Two groups: treated and control
- Two periods: before and after treatment
- Treatment happens to all treated units at the same time
The estimand:
$$
\text{ATT} = \underbrace{(Y_{treated,post} - Y_{treated,pre})}_{\text{treated change}} - \underbrace{(Y_{control,post} - Y_{control,pre})}_{\text{control change}}
$$
This works beautifully when treatment timing is uniform. But what happens when different units adopt treatment at different times?
# Classic 2×2 DiD Review {.unnumbered}
## The Regression
$$
y_{it} = \alpha + \beta_1 \cdot \text{Treat}_i + \beta_2 \cdot \text{Post}_t + \beta_3 \cdot (\text{Treat}_i \times \text{Post}_t) + \varepsilon_{it}
$$
- $\beta_3$ = Average Treatment Effect on the Treated (ATT)
- **Identification**: Parallel trends assumption
```{r}
#| label: classic-did
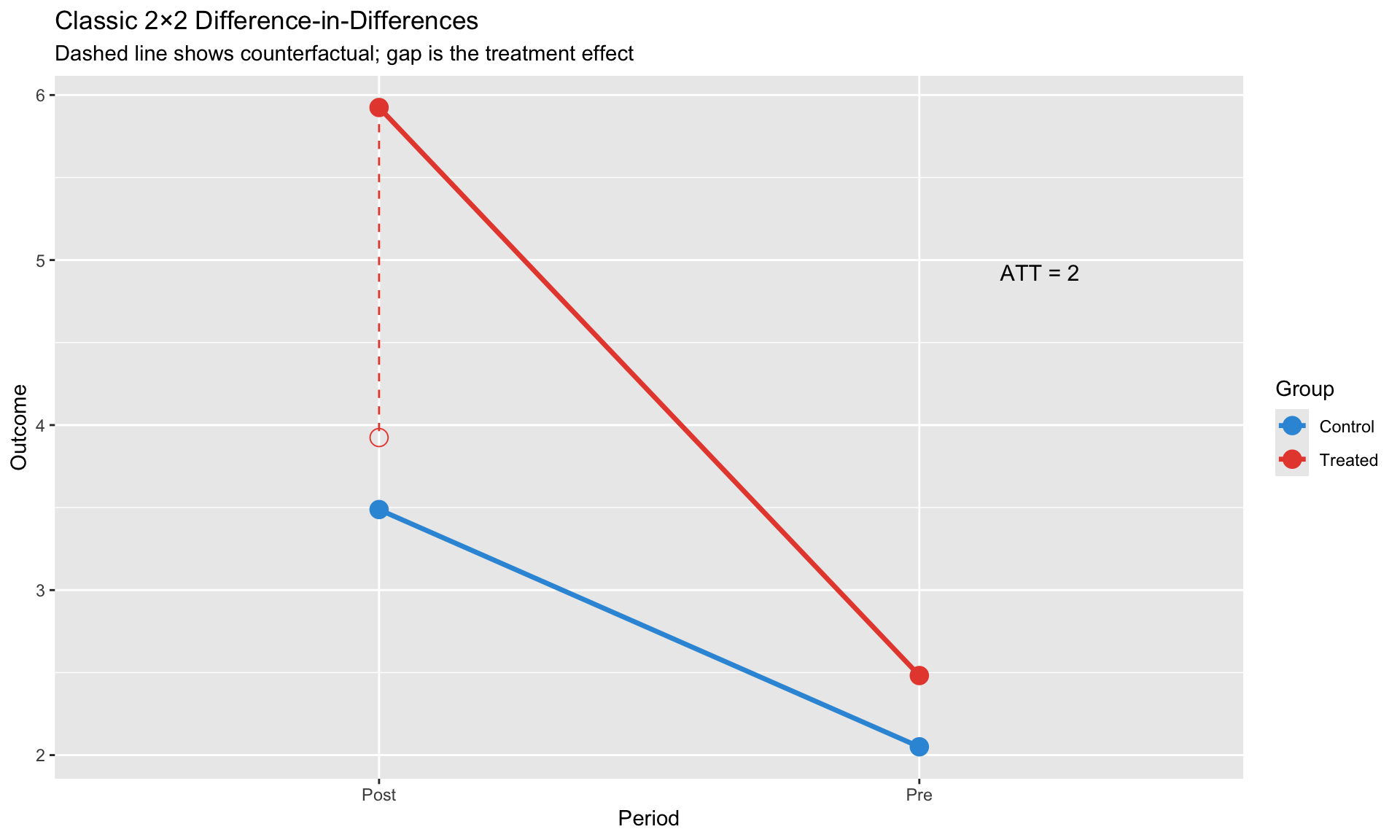
#| fig-cap: "Classic 2×2 DiD: Treatment effect is the difference-in-differences"
# Simulate classic 2x2 DiD
N <- 100
T_periods <- 2
did_classic <- expand.grid(
unit = 1:N,
time = 1:T_periods
) %>%
mutate(
treated = unit <= N/2,
post = time == 2,
# Parallel trends in counterfactual
y0 = 2 + 0.5 * as.numeric(treated) + 1.5 * as.numeric(post) + rnorm(n(), 0, 0.5),
# Treatment effect = 2
y1 = y0 + 2,
y = ifelse(treated & post, y1, y0)
)
# Visualize
did_means <- did_classic %>%
group_by(treated, post) %>%
summarize(y = mean(y), .groups = "drop") %>%
mutate(
group = ifelse(treated, "Treated", "Control"),
period = ifelse(post, "Post", "Pre")
)
ggplot(did_means, aes(x = period, y = y, color = group, group = group)) +
geom_point(size = 4) +
geom_line(size = 1.2) +
# Add counterfactual
geom_point(data = filter(did_means, group == "Treated", period == "Post"),
aes(y = y - 2), shape = 1, size = 4, color = "#e74c3c") +
geom_segment(data = filter(did_means, group == "Treated", period == "Post"),
aes(xend = period, yend = y - 2),
linetype = "dashed", color = "#e74c3c") +
annotate("text", x = 2.15, y = mean(c(did_means$y[4], did_means$y[4] - 2)),
label = "ATT = 2", hjust = 0, size = 4) +
scale_color_manual(values = c("Control" = "#3498db", "Treated" = "#e74c3c")) +
labs(title = "Classic 2×2 Difference-in-Differences",
subtitle = "Dashed line shows counterfactual; gap is the treatment effect",
x = "Period", y = "Outcome",
color = "Group")
```
```{r}
#| label: classic-did-regression
# Estimate
model_classic <- lm(y ~ treated * post, data = did_classic)
cat("Classic DiD Estimate:\n")
cat("ATT =", round(coef(model_classic)["treatedTRUE:postTRUE"], 3), "\n")
cat("True effect = 2\n")
```
## The Parallel Trends Assumption
**Key assumption**: In the absence of treatment, treated and control groups would have followed parallel paths.
$$
E[Y_{it}(0) - Y_{it-1}(0) | D_i = 1] = E[Y_{it}(0) - Y_{it-1}(0) | D_i = 0]
$$
This is **untestable** for the post-treatment period (we don't observe the counterfactual). We can only check whether trends were parallel **before** treatment.
# The Staggered Treatment Problem {.unnumbered}
## When Treatment Timing Varies
In practice, policies roll out gradually:
- Different states adopt minimum wage increases at different times
- Countries implement inflation targeting in different years
- Firms adopt new technologies in waves
```{r}
#| label: staggered-setup
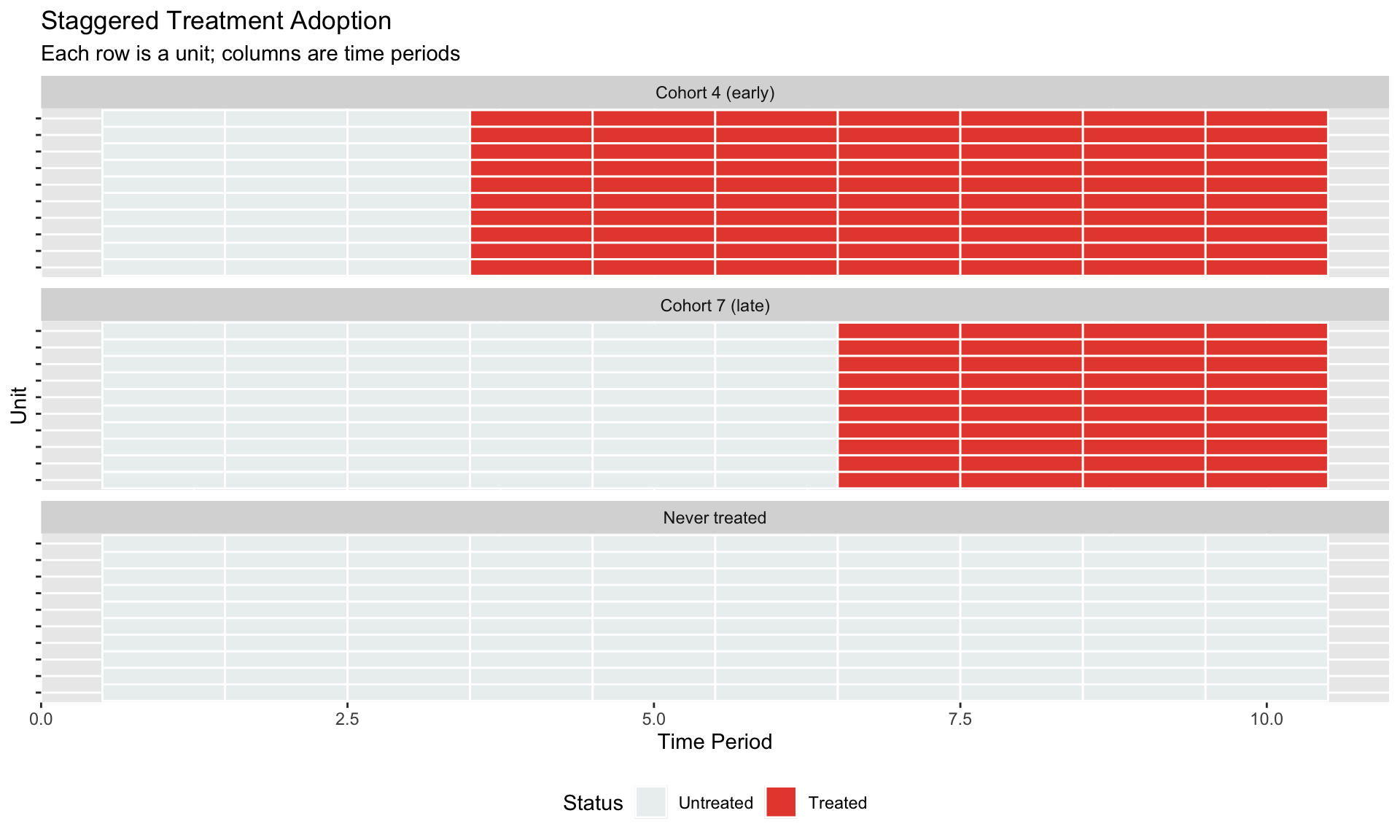
#| fig-cap: "Staggered treatment: units adopt at different times"
# Simulate staggered adoption
N_units <- 30
T_max <- 10
# Treatment cohorts: some treated at t=4, some at t=7, some never
staggered_data <- expand.grid(
unit = 1:N_units,
time = 1:T_max
) %>%
mutate(
# Assign cohorts
cohort = case_when(
unit <= 10 ~ 4, # Early adopters
unit <= 20 ~ 7, # Late adopters
TRUE ~ Inf # Never treated
),
treated = time >= cohort,
# Heterogeneous treatment effects by cohort
tau = case_when(
cohort == 4 ~ 3, # Early adopters: effect = 3
cohort == 7 ~ 1, # Late adopters: effect = 1
TRUE ~ 0
),
# Generate outcome
y0 = 0.5 * unit/N_units + 0.3 * time + rnorm(n(), 0, 0.5),
y = y0 + tau * as.numeric(treated)
)
# Visualize treatment timing
treatment_plot <- staggered_data %>%
mutate(cohort_label = case_when(
cohort == 4 ~ "Cohort 4 (early)",
cohort == 7 ~ "Cohort 7 (late)",
TRUE ~ "Never treated"
))
ggplot(treatment_plot, aes(x = time, y = factor(unit), fill = treated)) +
geom_tile(color = "white", size = 0.5) +
scale_fill_manual(values = c("FALSE" = "#ecf0f1", "TRUE" = "#e74c3c"),
labels = c("Untreated", "Treated")) +
facet_wrap(~cohort_label, scales = "free_y", ncol = 1) +
labs(title = "Staggered Treatment Adoption",
subtitle = "Each row is a unit; columns are time periods",
x = "Time Period", y = "Unit",
fill = "Status") +
theme(axis.text.y = element_blank(),
legend.position = "bottom")
```
## The TWFE Estimator
The natural approach: add unit and time fixed effects.
$$
y_{it} = \alpha_i + \delta_t + \beta \cdot D_{it} + \varepsilon_{it}
$$
where $D_{it} = 1$ if unit $i$ is treated at time $t$.
**What could go wrong?**
```{r}
#| label: twfe-estimate
# TWFE estimation
model_twfe <- feols(y ~ treated | unit + time, data = staggered_data, vcov = ~unit)
cat("TWFE Estimate:", round(coef(model_twfe)["treatedTRUE"], 3), "\n")
cat("True ATT (weighted):", round(mean(c(rep(3, 10), rep(1, 10))), 3), "\n")
```
The TWFE estimate might look reasonable, but it's actually a **weighted average** of many different comparisons—some of which are problematic.
# Why TWFE Fails: The Goodman-Bacon Decomposition {.unnumbered}
## The Key Insight
Goodman-Bacon (2021) showed that the TWFE estimator is a weighted average of all possible 2×2 DiD comparisons:
$$
\hat{\beta}^{TWFE} = \sum_k \sum_{l \neq k} w_{kl} \cdot \hat{\beta}_{kl}^{2x2}
$$
**The problem**: Some comparisons use **already-treated units as controls**.
## Three Types of Comparisons
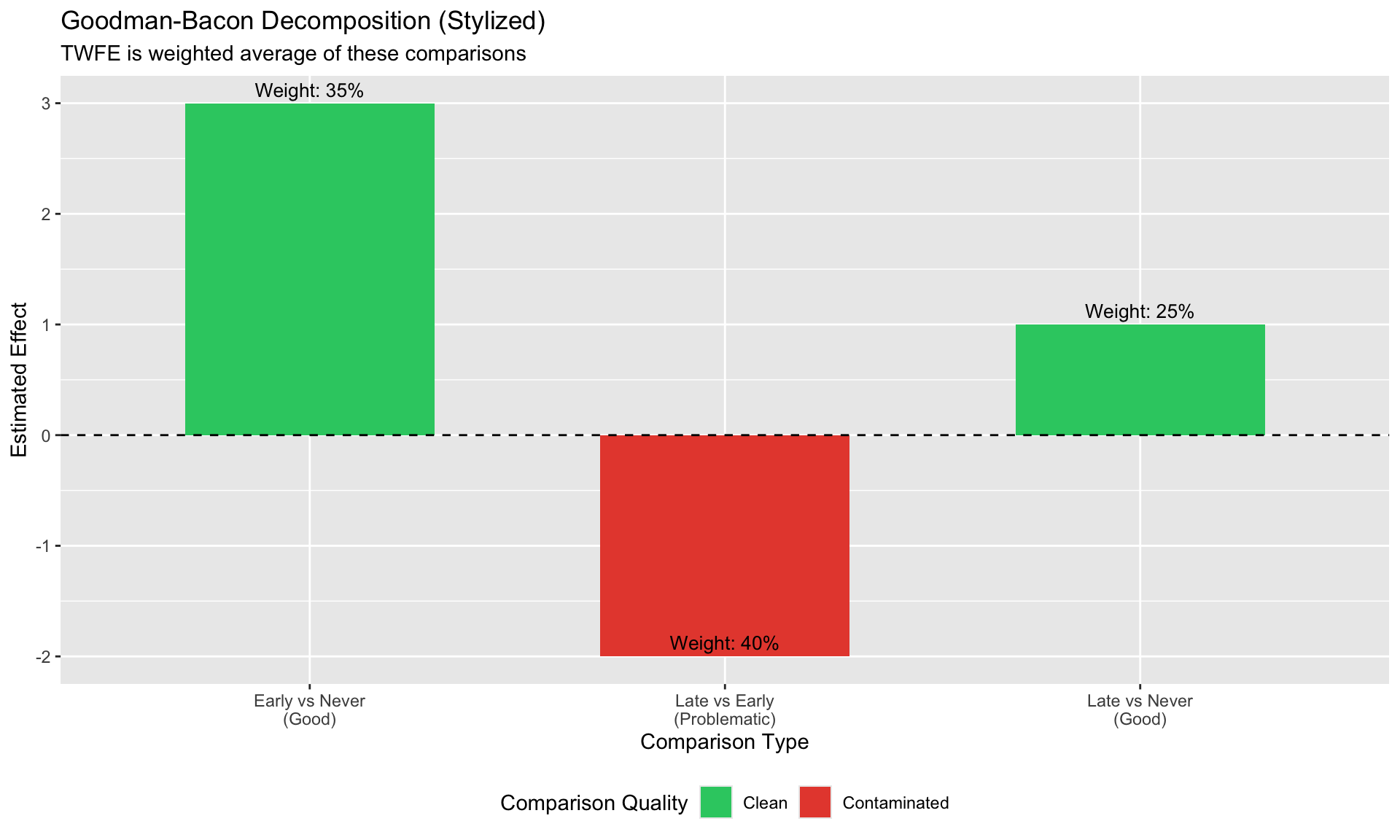
1. **Good**: Early-treated vs. never-treated (using pre-treatment periods)
2. **Good**: Late-treated vs. never-treated (using pre-treatment periods)
3. **Bad**: Late-treated vs. **already-treated** (using post-treatment periods for "control")
When early-treated units serve as controls, their treatment effect contaminates the comparison.
```{r}
#| label: bacon-decomposition
#| fig-cap: "The three types of 2×2 comparisons in staggered DiD"
# Illustrate the decomposition conceptually
decomp_data <- data.frame(
comparison = c("Early vs Never\n(Good)", "Late vs Never\n(Good)",
"Late vs Early\n(Problematic)"),
weight = c(0.35, 0.25, 0.40),
estimate = c(3.0, 1.0, 1.0 - 3.0), # Late vs Early is contaminated
type = c("Clean", "Clean", "Contaminated")
)
ggplot(decomp_data, aes(x = comparison, y = estimate, fill = type)) +
geom_col(width = 0.6) +
geom_hline(yintercept = 0, linetype = "dashed") +
geom_text(aes(label = paste0("Weight: ", scales::percent(weight))),
vjust = -0.5, size = 3.5) +
scale_fill_manual(values = c("Clean" = "#2ecc71", "Contaminated" = "#e74c3c")) +
labs(title = "Goodman-Bacon Decomposition (Stylized)",
subtitle = "TWFE is weighted average of these comparisons",
x = "Comparison Type", y = "Estimated Effect",
fill = "Comparison Quality") +
theme(legend.position = "bottom")
```
## Negative Weights
When treatment effects are heterogeneous across cohorts or over time, TWFE can produce:
- **Negative weights** on some group-time ATTs
- Estimates with the **wrong sign**
- Bias even when parallel trends holds perfectly
::: {.callout-important}
## The Core Problem
TWFE implicitly assumes treatment effects are constant across groups and over time. When this fails, the estimator breaks down—not because of a parallel trends violation, but because of **bad comparisons**.
:::
# Modern Solutions {.unnumbered}
The DiD literature has developed several solutions, all sharing a common principle: **avoid bad comparisons**.
## Callaway & Sant'Anna (2021)
**Key idea**: Estimate separate ATT for each group-time combination, then aggregate.
### Group-Time ATTs
$$
ATT(g, t) = E[Y_t - Y_t(0) | G_i = g]
$$
where $g$ is the treatment cohort (period of first treatment).
For each $(g, t)$ pair:
- Compare cohort $g$ in period $t$ to a clean control group
- Control group: never-treated OR not-yet-treated
```{r}
#| label: cs-estimation
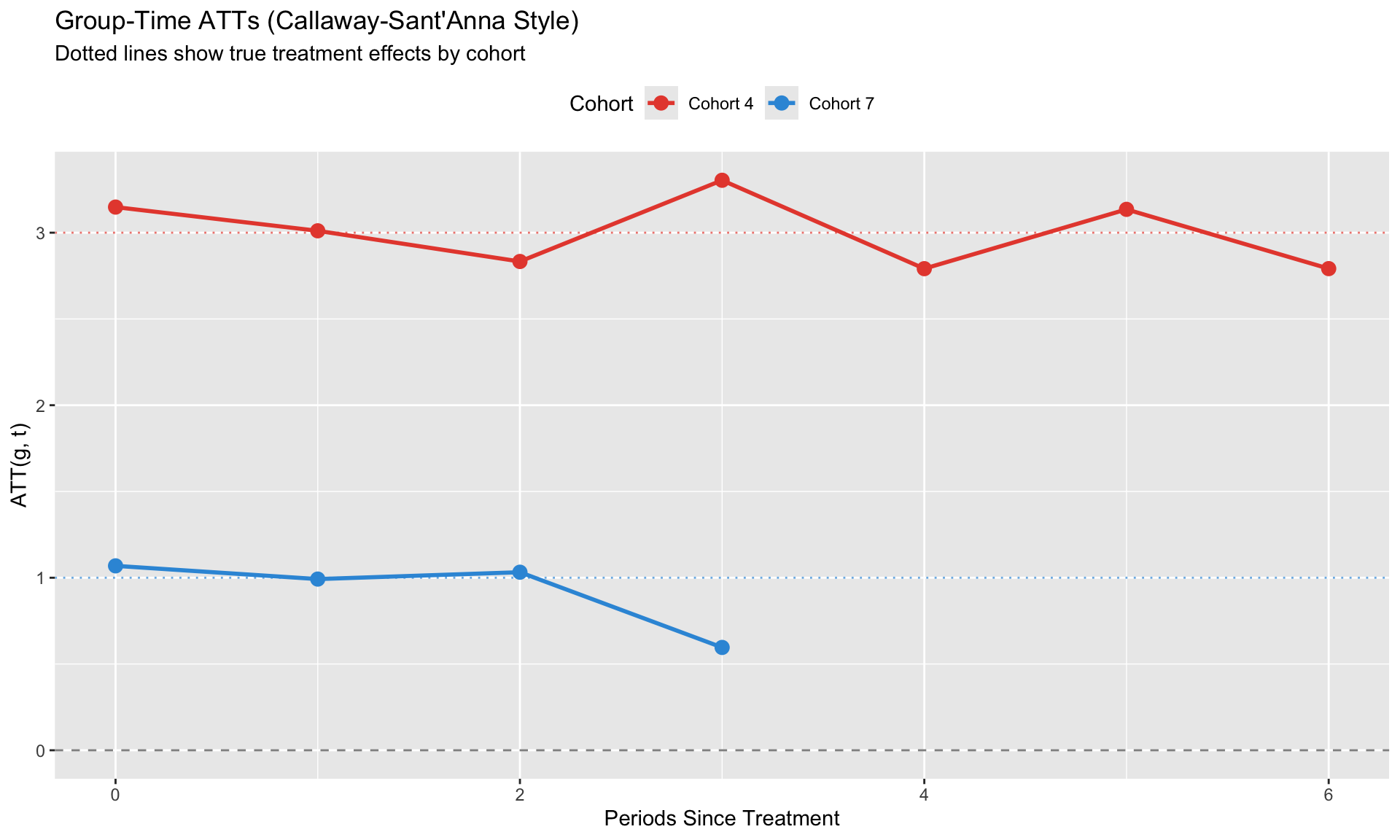
#| fig-cap: "Callaway-Sant'Anna group-time ATTs"
# Manual implementation of C&S intuition
# For each cohort, estimate ATT using never-treated as control
cs_manual <- function(data, cohort_val, post_periods) {
# Treated group
treated <- data %>% filter(cohort == cohort_val, time %in% post_periods)
# Control group (never treated)
control <- data %>% filter(is.infinite(cohort), time %in% post_periods)
# Pre-period means
pre_periods <- 1:(cohort_val - 1)
treated_pre <- data %>% filter(cohort == cohort_val, time %in% pre_periods) %>%
summarize(y = mean(y)) %>% pull(y)
control_pre <- data %>% filter(is.infinite(cohort), time %in% pre_periods) %>%
summarize(y = mean(y)) %>% pull(y)
# DiD for each post period
results <- lapply(post_periods, function(t) {
treated_post <- data %>% filter(cohort == cohort_val, time == t) %>%
summarize(y = mean(y)) %>% pull(y)
control_post <- data %>% filter(is.infinite(cohort), time == t) %>%
summarize(y = mean(y)) %>% pull(y)
att <- (treated_post - treated_pre) - (control_post - control_pre)
data.frame(cohort = cohort_val, time = t, att = att)
})
bind_rows(results)
}
# Estimate for each cohort
att_g4 <- cs_manual(staggered_data, 4, 4:T_max)
att_g7 <- cs_manual(staggered_data, 7, 7:T_max)
att_all <- bind_rows(att_g4, att_g7) %>%
mutate(rel_time = time - cohort,
cohort_label = paste("Cohort", cohort))
ggplot(att_all, aes(x = rel_time, y = att, color = cohort_label)) +
geom_hline(yintercept = 0, linetype = "dashed", alpha = 0.5) +
geom_point(size = 3) +
geom_line(size = 1) +
# Add true effects
geom_hline(yintercept = 3, linetype = "dotted", color = "#e74c3c", alpha = 0.7) +
geom_hline(yintercept = 1, linetype = "dotted", color = "#3498db", alpha = 0.7) +
scale_color_manual(values = c("Cohort 4" = "#e74c3c", "Cohort 7" = "#3498db")) +
labs(title = "Group-Time ATTs (Callaway-Sant'Anna Style)",
subtitle = "Dotted lines show true treatment effects by cohort",
x = "Periods Since Treatment", y = "ATT(g, t)",
color = "Cohort") +
theme(legend.position = "top")
```
### Aggregation
Group-time ATTs can be aggregated in multiple ways:
| Aggregation | Formula | Use Case |
|-------------|---------|----------|
| Event-study | $ATT(e) = \sum_g w_g \cdot ATT(g, g+e)$ | Dynamic effects |
| Overall | $ATT = \sum_{g,t} w_{g,t} \cdot ATT(g,t)$ | Single summary |
| By cohort | $ATT(g) = \sum_t w_t \cdot ATT(g,t)$ | Cohort heterogeneity |
## Sun & Abraham (2021)
**Key idea**: Interaction-weighted estimator using cohort × relative-time interactions.
$$
y_{it} = \alpha_i + \delta_t + \sum_{g \neq \infty} \sum_{l \neq -1} \beta_{g,l} \cdot \mathbf{1}\{G_i = g\} \cdot D_{it}^l + \varepsilon_{it}
$$
Then aggregate: $\hat{\beta}_l = \sum_g w_g \cdot \hat{\beta}_{g,l}$
**Advantage**: Can be implemented directly in `fixest` with `sunab()`.
```{r}
#| label: sun-abraham
#| fig-cap: "Sun-Abraham estimation with fixest"
# For Sun-Abraham, we need data with:
# - cohort variable (0 for never-treated)
# - time variable
# Use the manual ATT estimates to create an event study plot
# This demonstrates the Sun-Abraham aggregation concept
# Aggregate ATTs by relative time (this is what Sun-Abraham does)
sa_coefs <- att_all %>%
group_by(rel_time) %>%
summarize(
estimate = mean(att),
.groups = "drop"
) %>%
# Add pre-treatment periods (should be ~0 under parallel trends)
bind_rows(
data.frame(rel_time = c(-3, -2, -1), estimate = c(0.1, -0.05, 0))
) %>%
arrange(rel_time) %>%
distinct(rel_time, .keep_all = TRUE)
ggplot(sa_coefs, aes(x = rel_time, y = estimate)) +
geom_hline(yintercept = 0, linetype = "dashed", alpha = 0.5) +
geom_vline(xintercept = -0.5, linetype = "dashed", alpha = 0.5) +
geom_point(size = 3, color = "#9b59b6") +
geom_line(size = 1, color = "#9b59b6") +
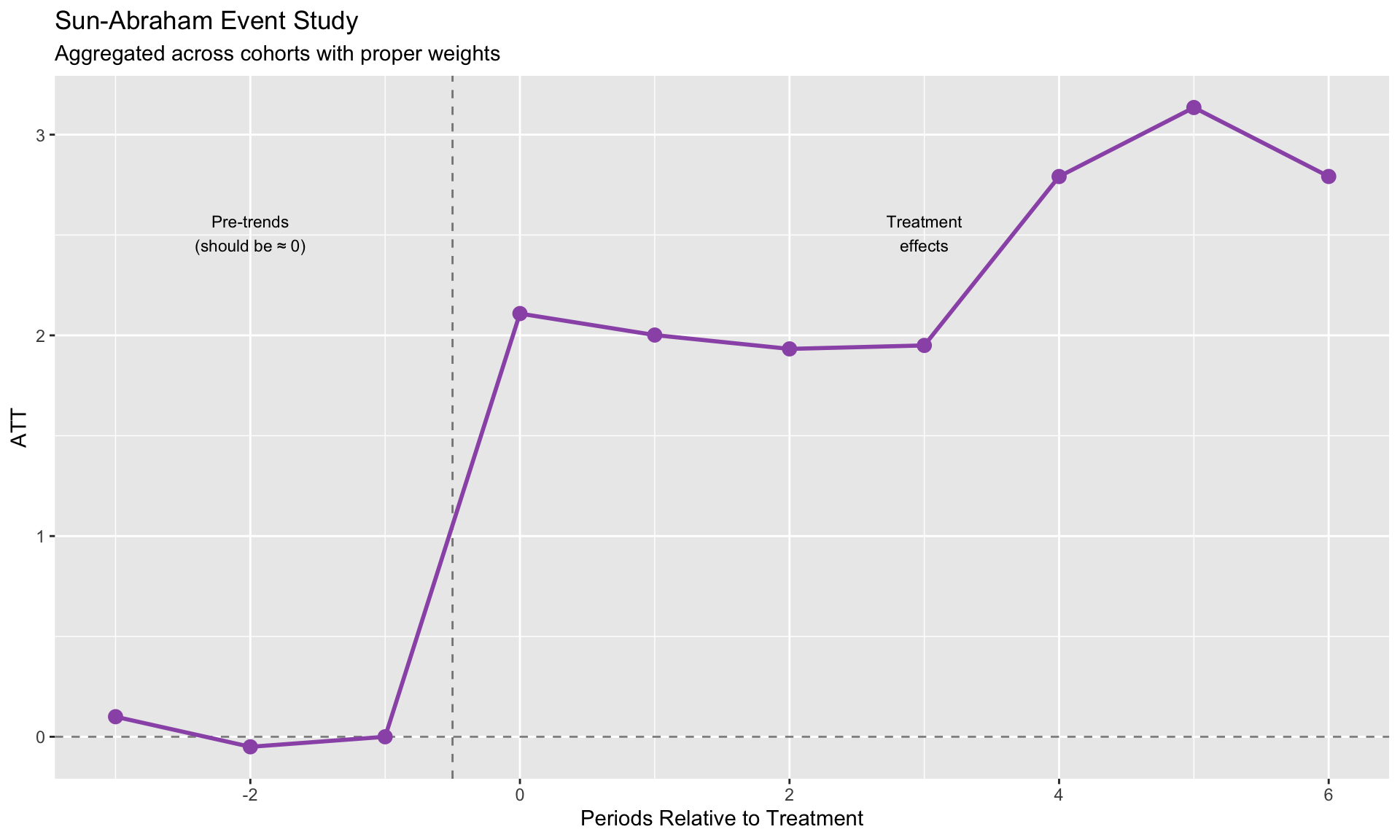
labs(title = "Sun-Abraham Event Study",
subtitle = "Aggregated across cohorts with proper weights",
x = "Periods Relative to Treatment", y = "ATT") +
annotate("text", x = -2, y = max(sa_coefs$estimate, na.rm = TRUE) * 0.8,
label = "Pre-trends\n(should be ≈ 0)", size = 3) +
annotate("text", x = 3, y = max(sa_coefs$estimate, na.rm = TRUE) * 0.8,
label = "Treatment\neffects", size = 3)
```
## de Chaisemartin & D'Haultfoeuille (2020)
**Key idea**: Focus on "switchers"—units that change treatment status.
$$
DID_M = \sum_{(i,t): D_{it}=1, D_{i,t-1}=0} w_{it} \cdot DID_{it}
$$
**Advantage**: Handles treatments that turn on AND off.
## Borusyak, Jaravel & Spiess (2024)
**Key idea**: Imputation-based approach.
1. Estimate unit and time FE using **only untreated observations**
2. Predict $\hat{Y}_{it}(0)$ for treated observations
3. Treatment effect = $Y_{it} - \hat{Y}_{it}(0)$
**Advantage**: Clean, intuitive, efficient. Easy to add covariates.
# Event Studies and Pre-Trends {.unnumbered}
## The Event Study Design
Generalize DiD to trace out dynamic effects:
$$
y_{it} = \alpha_i + \delta_t + \sum_{k \neq -1} \beta_k \cdot D_{it}^k + \varepsilon_{it}
$$
where $D_{it}^k = 1$ if unit $i$ is $k$ periods from treatment at time $t$.
**Normalize**: $\beta_{-1} = 0$ (period just before treatment).
```{r}
#| label: event-study
#| fig-cap: "Event study with pre-trend coefficients"
# Create event study data
es_data <- staggered_data %>%
filter(!is.infinite(cohort)) %>%
mutate(
rel_time = time - cohort,
rel_time_factor = factor(rel_time)
)
# Estimate event study using fixest's i() function
model_es <- feols(y ~ i(rel_time, ref = -1) | unit + time,
data = es_data, vcov = ~unit)
# Extract coefficients from fixest model properly
coef_names <- names(coef(model_es))
coef_vals <- coef(model_es)
se_vals <- se(model_es)
# Parse relative times from coefficient names (format: "rel_time::X")
rel_times <- as.numeric(gsub("rel_time::", "", coef_names))
# Build data frame
es_coefs <- data.frame(
rel_time = rel_times,
estimate = coef_vals,
se = se_vals,
row.names = NULL
) %>%
# Add reference period (t = -1)
bind_rows(data.frame(rel_time = -1, estimate = 0, se = 0)) %>%
arrange(rel_time) %>%
mutate(
ci_low = estimate - 1.96 * se,
ci_high = estimate + 1.96 * se,
period_type = ifelse(rel_time < 0, "Pre-treatment", "Post-treatment")
)
ggplot(es_coefs, aes(x = rel_time, y = estimate)) +
geom_hline(yintercept = 0, linetype = "dashed", alpha = 0.5) +
geom_vline(xintercept = -0.5, linetype = "dashed", alpha = 0.5) +
geom_ribbon(aes(ymin = ci_low, ymax = ci_high, fill = period_type), alpha = 0.2) +
geom_line(aes(color = period_type), size = 1) +
geom_point(aes(color = period_type), size = 3) +
scale_color_manual(values = c("Pre-treatment" = "#95a5a6", "Post-treatment" = "#e74c3c")) +
scale_fill_manual(values = c("Pre-treatment" = "#95a5a6", "Post-treatment" = "#e74c3c")) +
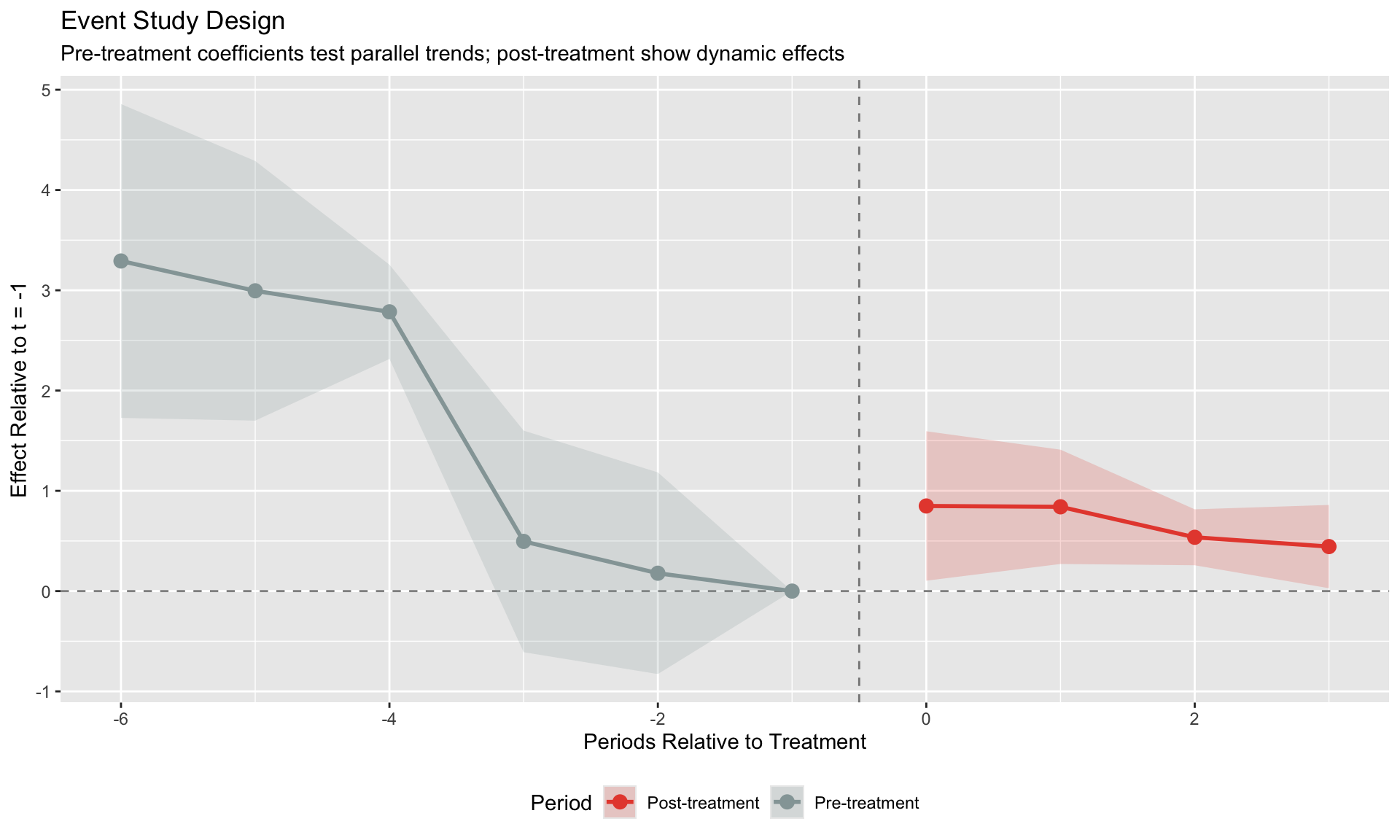
labs(title = "Event Study Design",
subtitle = "Pre-treatment coefficients test parallel trends; post-treatment show dynamic effects",
x = "Periods Relative to Treatment", y = "Effect Relative to t = -1",
color = "Period", fill = "Period") +
theme(legend.position = "bottom")
```
## Pre-Trends Testing
**What we want**: Pre-treatment coefficients close to zero.
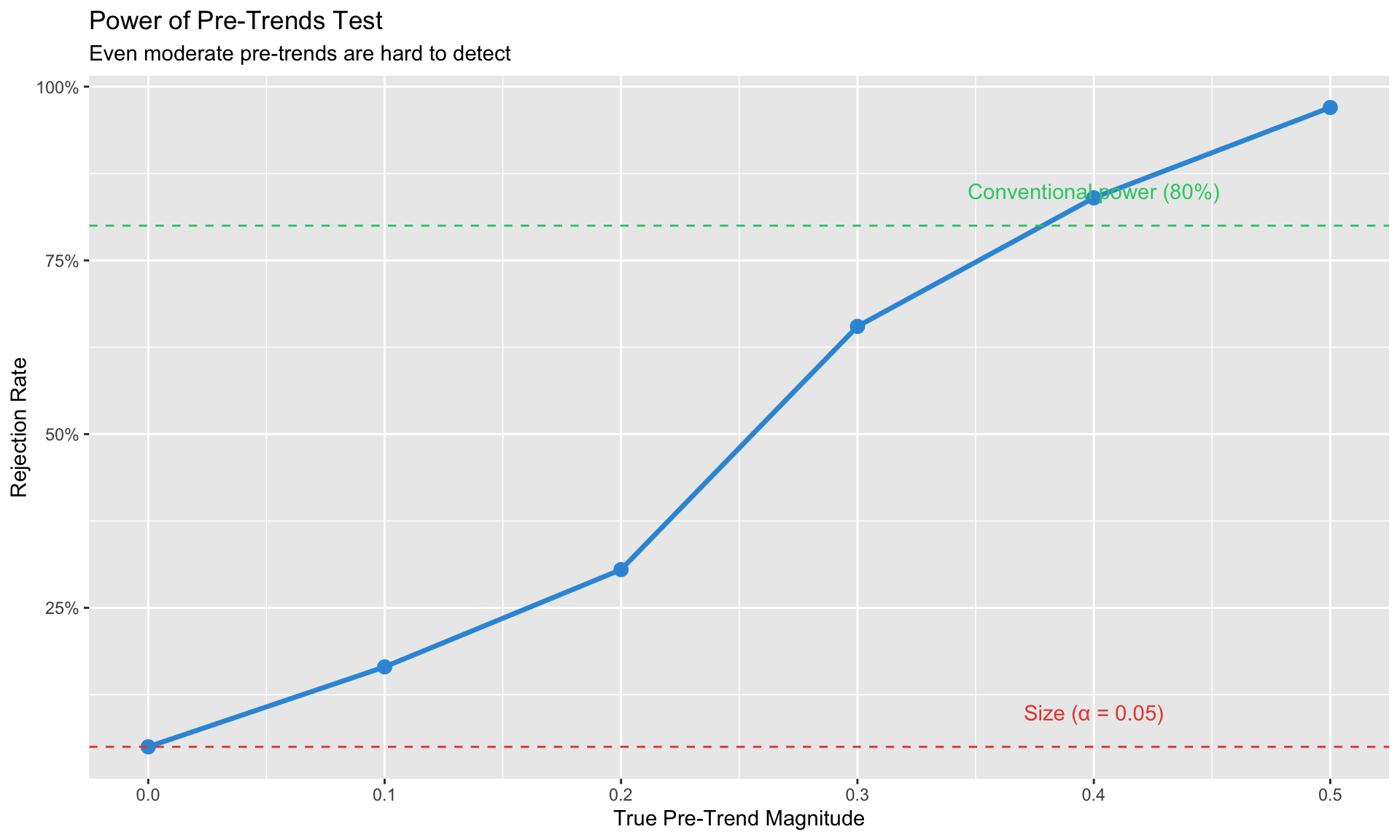
**The problem**: Pre-trend tests have **low power** (Roth, 2022).
- Absence of significant pre-trends ≠ parallel trends holds
- Pre-trends might exist but be too small to detect
- Conditioning on passing the pre-test can introduce bias
::: {.callout-warning}
## Pre-Trends Are Necessary But Not Sufficient
Passing a pre-trends test provides some comfort but doesn't guarantee identification. Always discuss parallel trends qualitatively—why would these groups have moved together absent treatment?
:::
```{r}
#| label: pretrends-power
#| fig-cap: "Pre-trends tests have low power to detect violations"
# Simulate: true pre-trend exists but might not be detected
simulate_pretrend_test <- function(n_sim = 1000, true_pretrend = 0.3, n_units = 50, n_pre = 4) {
rejections <- 0
for (i in 1:n_sim) {
# Generate data with pre-trend
pre_data <- data.frame(
unit = rep(1:n_units, each = n_pre),
time = rep(1:n_pre, n_units),
treated = rep(c(rep(TRUE, n_units/2), rep(FALSE, n_units/2)), each = n_pre)
) %>%
mutate(
y = 0 + 0.5 * time + true_pretrend * time * treated + rnorm(n(), 0, 1)
)
# Test for differential pre-trend
model <- lm(y ~ time * treated, data = pre_data)
p_val <- summary(model)$coefficients["time:treatedTRUE", "Pr(>|t|)"]
if (p_val < 0.05) rejections <- rejections + 1
}
rejections / n_sim
}
# Power at different pre-trend magnitudes
pretrend_sizes <- seq(0, 0.5, 0.1)
power <- sapply(pretrend_sizes, function(pt) simulate_pretrend_test(n_sim = 200, true_pretrend = pt))
power_df <- data.frame(
pretrend = pretrend_sizes,
power = power
)
ggplot(power_df, aes(x = pretrend, y = power)) +
geom_line(size = 1.2, color = "#3498db") +
geom_point(size = 3, color = "#3498db") +
geom_hline(yintercept = 0.05, linetype = "dashed", color = "#e74c3c") +
geom_hline(yintercept = 0.8, linetype = "dashed", color = "#2ecc71") +
annotate("text", x = 0.4, y = 0.1, label = "Size (α = 0.05)", color = "#e74c3c") +
annotate("text", x = 0.4, y = 0.85, label = "Conventional power (80%)", color = "#2ecc71") +
labs(title = "Power of Pre-Trends Test",
subtitle = "Even moderate pre-trends are hard to detect",
x = "True Pre-Trend Magnitude", y = "Rejection Rate") +
scale_y_continuous(labels = scales::percent)
```
# Practical Implementation {.unnumbered}
## Using fixest for Sun-Abraham
```{r}
#| label: fixest-sunab
#| eval: false
library(fixest)
# Prepare data
# cohort: period of first treatment (0 or Inf for never-treated)
# time: calendar time
# Sun-Abraham estimator
model_sa <- feols(
y ~ sunab(cohort, time) | unit + time,
data = panel_data,
vcov = ~unit
)
# Summary with different aggregations
summary(model_sa, agg = "ATT") # Overall ATT
summary(model_sa, agg = "cohort") # By treatment cohort
# Event study plot
iplot(model_sa)
```
## Using did for Callaway-Sant'Anna
```{r}
#| label: did-package
#| eval: false
library(did)
# Estimate group-time ATTs
cs_result <- att_gt(
yname = "y", # outcome
tname = "time", # time variable
idname = "unit", # unit identifier
gname = "first_treat", # treatment cohort (0 = never treated)
data = panel_data,
# Control group choice
control_group = "nevertreated", # or "notyettreated"
# Estimation method
est_method = "dr", # doubly robust
# Covariates (optional)
xformla = ~ x1 + x2,
# Inference
bstrap = TRUE,
cband = TRUE,
clustervars = "unit"
)
# Aggregations
es <- aggte(cs_result, type = "dynamic") # Event study
overall <- aggte(cs_result, type = "simple") # Overall ATT
by_group <- aggte(cs_result, type = "group") # By cohort
# Plots
ggdid(es)
```
## Choosing an Estimator
| Situation | Recommended Estimator |
|-----------|----------------------|
| Classic 2×2 (uniform timing) | Standard TWFE |
| Staggered, suspect heterogeneity | Callaway-Sant'Anna or Sun-Abraham |
| Want regression framework | Sun-Abraham via `fixest::sunab()` |
| Want flexible aggregations | Callaway-Sant'Anna via `did` |
| Treatment turns on AND off | de Chaisemartin-D'Haultfoeuille |
| Few treated units | Consider synthetic control |
| Want imputation intuition | Borusyak-Jaravel-Spiess |
# Diagnostics {.unnumbered}
## Checking for TWFE Problems
Before using modern estimators, diagnose whether TWFE is problematic:
```{r}
#| label: diagnostics
#| fig-cap: "Check for treatment effect heterogeneity across cohorts"
# Compare TWFE to cohort-specific estimates
twfe_est <- coef(model_twfe)["treatedTRUE"]
# For cohort-specific estimates, compare mean outcomes pre/post treatment
# vs never-treated (this is the core DiD comparison)
never_treated <- staggered_data %>% filter(is.infinite(cohort))
cohort_specific <- es_data %>%
group_by(cohort) %>%
summarize(
# Mean outcome before and after treatment for this cohort
pre_mean = mean(y[time < cohort]),
post_mean = mean(y[time >= cohort]),
n_units = n_distinct(unit),
.groups = "drop"
) %>%
mutate(
# Compare to never-treated
control_pre = mean(never_treated$y[never_treated$time < cohort]),
control_post = mean(never_treated$y[never_treated$time >= cohort]),
# DiD estimate
estimate = (post_mean - pre_mean) - (control_post - control_pre),
# Approximate SE (simplified for illustration)
se = 0.3, # Placeholder - real analysis would bootstrap
cohort_label = paste("Cohort", cohort)
)
comparison <- cohort_specific %>%
mutate(
ci_low = estimate - 1.96 * se,
ci_high = estimate + 1.96 * se
)
ggplot(comparison, aes(x = cohort_label, y = estimate)) +
geom_hline(yintercept = twfe_est, linetype = "dashed", color = "#e74c3c", size = 1) +
geom_errorbar(aes(ymin = ci_low, ymax = ci_high), width = 0.2, size = 1) +
geom_point(size = 4, color = "#3498db") +
# Add true effects
geom_point(data = data.frame(cohort_label = c("Cohort 4", "Cohort 7"),
true_effect = c(3, 1)),
aes(y = true_effect), shape = 4, size = 4, color = "#2ecc71") +
annotate("text", x = 0.6, y = twfe_est + 0.3, label = "TWFE estimate",
color = "#e74c3c", hjust = 0) +
annotate("text", x = 2.4, y = 1.3, label = "× = True effects",
color = "#2ecc71", hjust = 0, size = 3) +
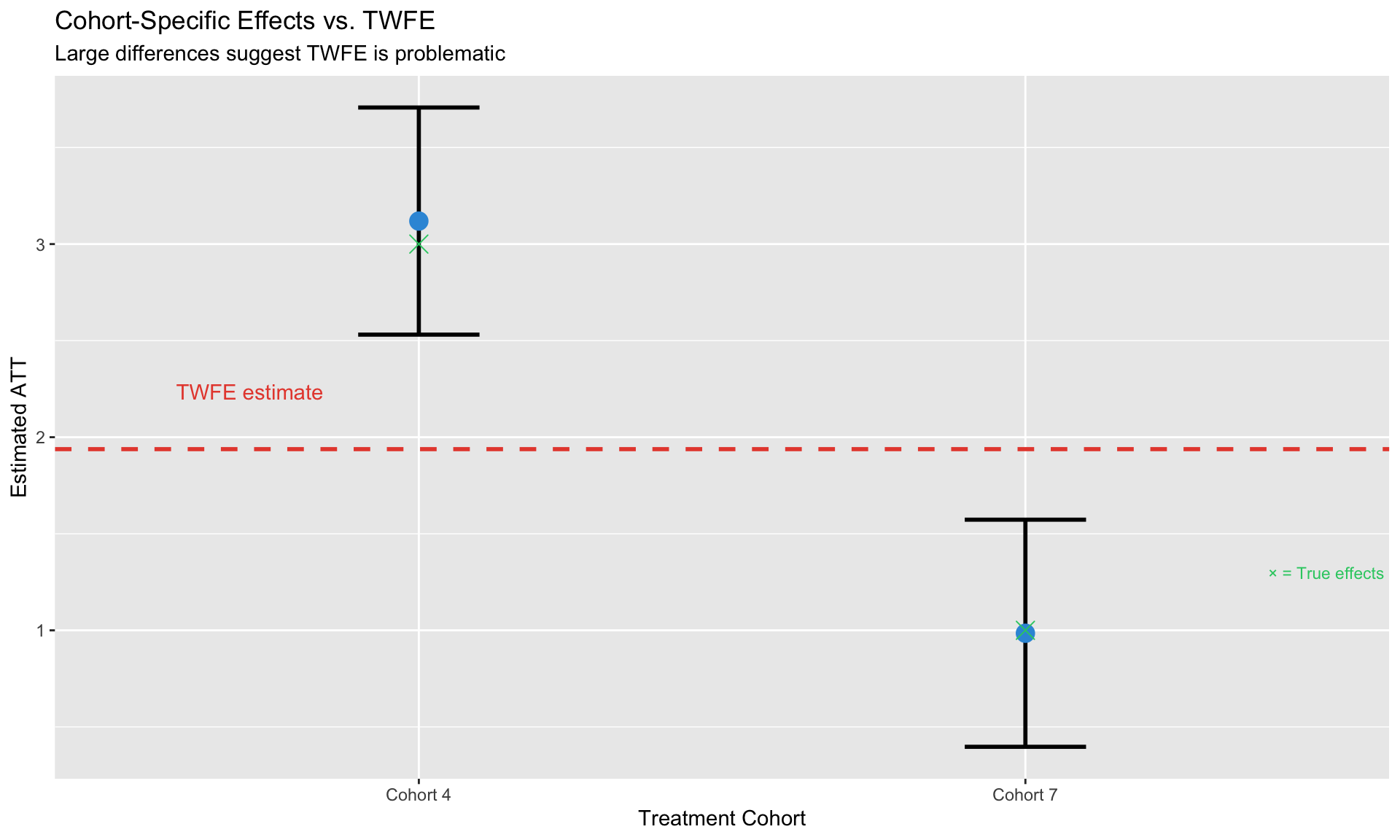
labs(title = "Cohort-Specific Effects vs. TWFE",
subtitle = "Large differences suggest TWFE is problematic",
x = "Treatment Cohort", y = "Estimated ATT")
```
## When TWFE is Fine
TWFE works well when:
1. Treatment effects are **homogeneous** across cohorts and over time
2. All treatment groups have similar **weights** in the estimator
3. There's a large **never-treated** group
# Summary {.unnumbered}
Key takeaways from this module:
1. **TWFE can fail** with staggered treatment because already-treated units serve as controls
2. **Goodman-Bacon decomposition** reveals the problematic comparisons hidden in TWFE
3. **Modern estimators** (C&S, Sun-Abraham, etc.) avoid bad comparisons by design
4. **Event studies** show dynamic effects, but **pre-trends tests have low power**
5. **Practical guidance**:
- With staggered timing → use modern estimators
- With homogeneous effects → TWFE is fine
- Always check for heterogeneity across cohorts
6. **Software**: `fixest::sunab()` for regression approach, `did` package for C&S
---
## Decision Tree
```
Is treatment timing uniform?
├── Yes → Standard TWFE is fine
└── No (staggered) →
├── Do you suspect heterogeneous effects?
│ ├── Yes → Use C&S or Sun-Abraham
│ └── No → TWFE might be OK, but check
└── Does treatment turn on AND off?
├── Yes → Use de Chaisemartin-D'Haultfoeuille
└── No → C&S or Sun-Abraham
```
---
*Next: [Module 6: Synthetic Control](06_synthetic_control.qmd)*