---
title: "Bayesian Foundations"
subtitle: "From Priors to Posteriors in Macroeconometrics"
---
```{r}
#| label: setup
#| include: false
knitr::opts_chunk$set(
echo = TRUE, warning = FALSE, message = FALSE,
fig.width = 10, fig.height = 6, fig.align = "center"
)
library(dplyr)
library(tidyr)
library(ggplot2)
theme_set(theme_minimal(base_size = 13))
set.seed(42)
```
# The Bayesian Paradigm {.unnumbered}
Bayesian inference provides a coherent framework for combining prior knowledge with data to quantify uncertainty about parameters. In macroeconometrics, where samples are small and prior information is often available, Bayesian methods have become essential.
## Bayes' Theorem
The foundation of Bayesian inference:
$$
\underbrace{p(\theta | Y)}_{\text{Posterior}} = \frac{\overbrace{p(Y | \theta)}^{\text{Likelihood}} \times \overbrace{p(\theta)}^{\text{Prior}}}{\underbrace{p(Y)}_{\text{Marginal Likelihood}}}
$$
Or, since $p(Y)$ is a constant given the data:
$$
p(\theta | Y) \propto p(Y | \theta) \times p(\theta)
$$
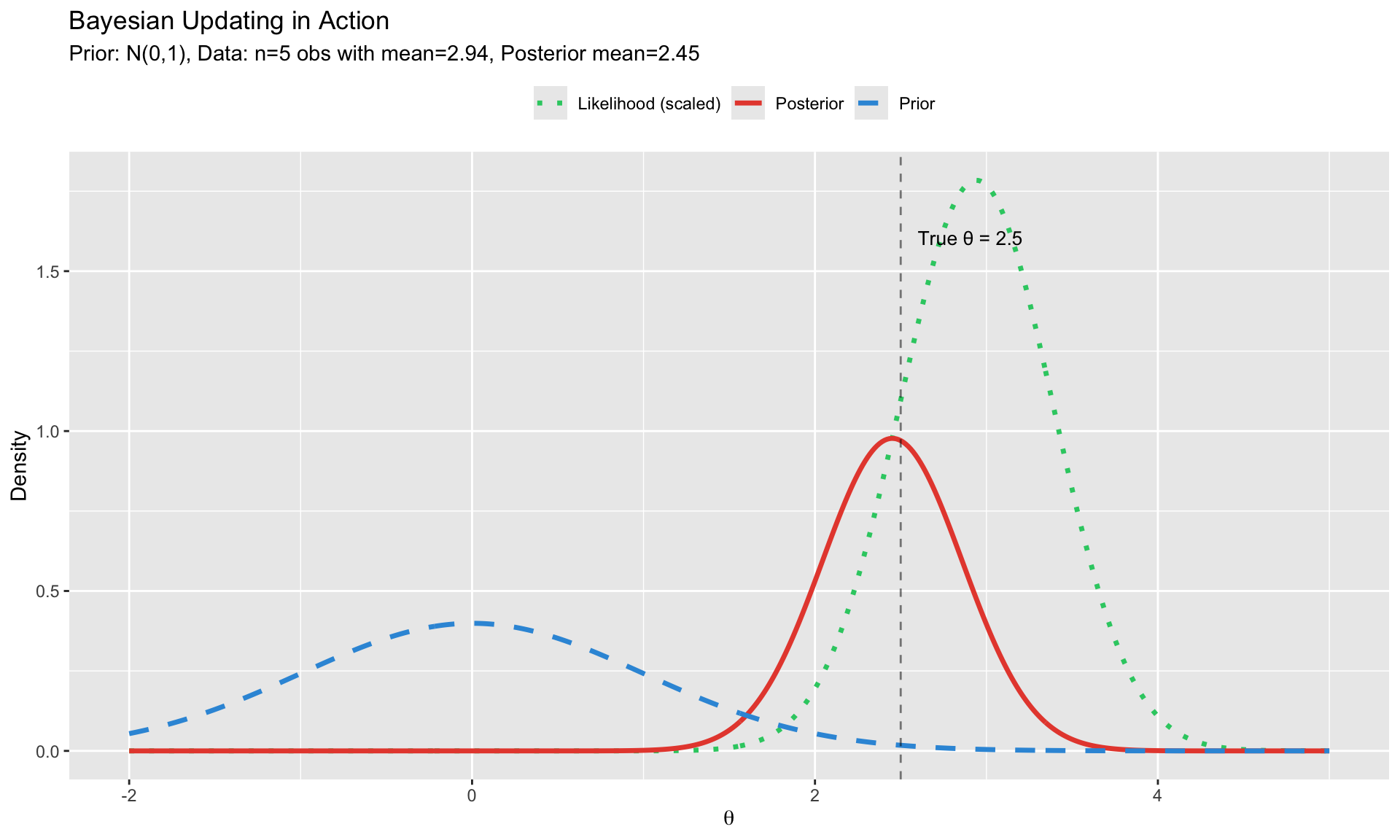
**In words**: What we believe after seeing the data (posterior) is proportional to what the data tells us (likelihood) times what we believed before (prior).
```{r}
#| label: bayes-visual
#| fig-cap: "Bayesian updating: Prior × Likelihood → Posterior"
# Demonstrate Bayesian updating
# Example: estimating a mean with known variance
# True parameter
theta_true <- 2.5
# Prior: N(0, 1) - vague prior centered at 0
prior_mean <- 0
prior_var <- 1
# Data: 5 observations with known variance
n <- 5
sigma2 <- 1 # known variance
y <- rnorm(n, theta_true, sqrt(sigma2))
y_bar <- mean(y)
# Posterior (conjugate update)
# For normal-normal: posterior_mean = weighted average
# posterior_var = 1/(1/prior_var + n/sigma2)
post_var <- 1 / (1/prior_var + n/sigma2)
post_mean <- post_var * (prior_mean/prior_var + n*y_bar/sigma2)
# Plot
theta_grid <- seq(-2, 5, length.out = 500)
plot_data <- data.frame(
theta = rep(theta_grid, 3),
density = c(
dnorm(theta_grid, prior_mean, sqrt(prior_var)),
dnorm(theta_grid, y_bar, sqrt(sigma2/n)) * 2, # scaled likelihood
dnorm(theta_grid, post_mean, sqrt(post_var))
),
type = rep(c("Prior", "Likelihood (scaled)", "Posterior"), each = length(theta_grid))
)
ggplot(plot_data, aes(x = theta, y = density, color = type, linetype = type)) +
geom_line(size = 1.2) +
geom_vline(xintercept = theta_true, linetype = "dashed", alpha = 0.5) +
annotate("text", x = theta_true + 0.1, y = max(plot_data$density) * 0.9,
label = paste0("True θ = ", theta_true), hjust = 0, size = 3.5) +
scale_color_manual(values = c("Prior" = "#3498db",
"Likelihood (scaled)" = "#2ecc71",
"Posterior" = "#e74c3c")) +
scale_linetype_manual(values = c("Prior" = "dashed",
"Likelihood (scaled)" = "dotted",
"Posterior" = "solid")) +
labs(title = "Bayesian Updating in Action",
subtitle = paste0("Prior: N(0,1), Data: n=", n, " obs with mean=",
round(y_bar, 2), ", Posterior mean=", round(post_mean, 2)),
x = expression(theta), y = "Density",
color = "", linetype = "") +
theme(legend.position = "top")
```
## Key Components
| Component | Symbol | Interpretation |
|-----------|--------|----------------|
| **Prior** | $p(\theta)$ | Belief about $\theta$ before seeing data |
| **Likelihood** | $p(Y|\theta)$ | Probability of data given parameters |
| **Posterior** | $p(\theta|Y)$ | Updated belief after seeing data |
| **Marginal Likelihood** | $p(Y)$ | Probability of data (for model comparison) |
# Why Bayesian for Macroeconometrics? {.unnumbered}
## Challenge 1: Small Samples
Macroeconomic data is scarce:
- Quarterly GDP: ~80 years = 320 observations
- Post-WWII monetary policy: ~75 years
- Post-Great Moderation: ~40 years
With many parameters and few observations, frequentist methods suffer from:
- Overfitting
- Imprecise estimates
- Unreliable asymptotic inference
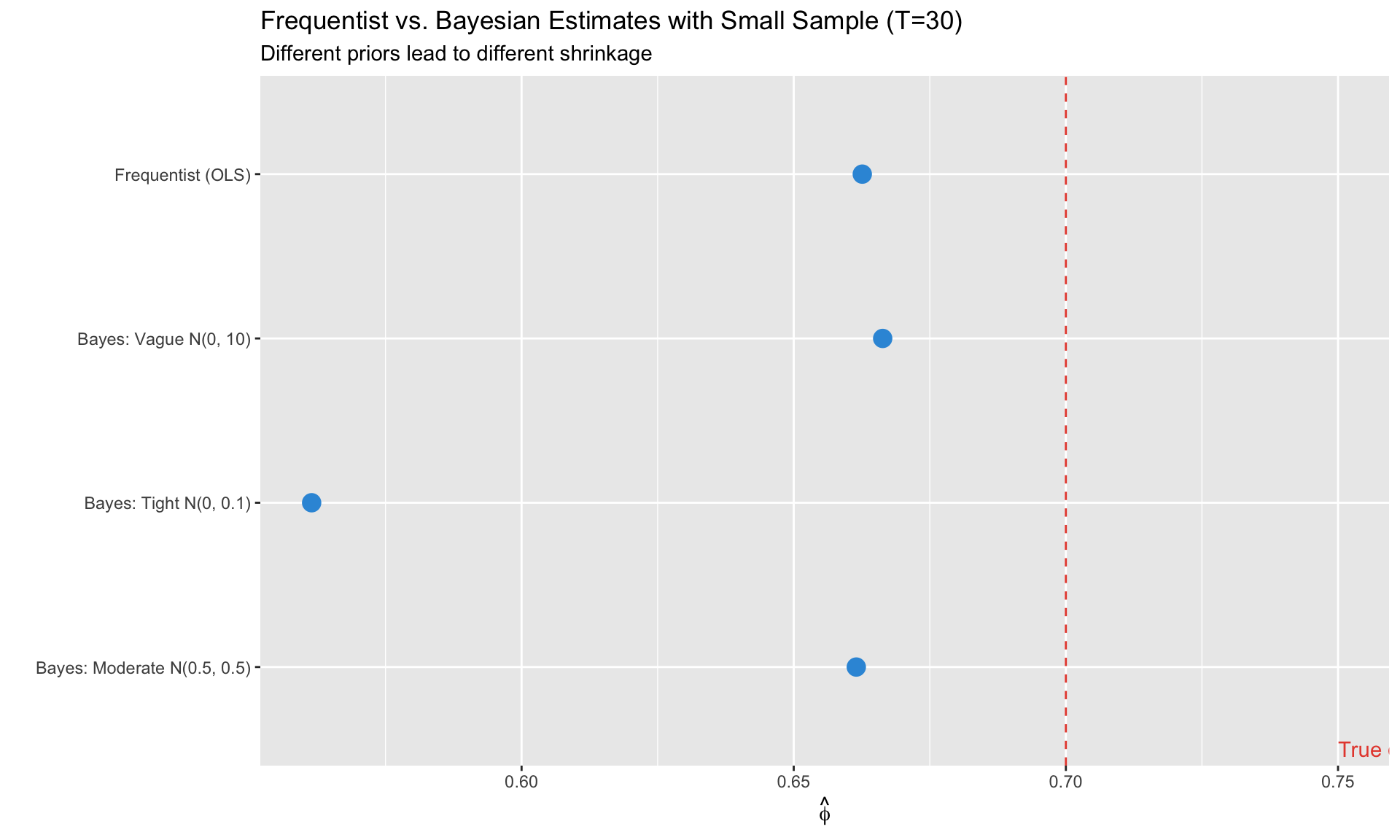
**Bayesian solution**: Priors regularize estimation, preventing overfitting by shrinking toward sensible values.
```{r}
#| label: small-sample
#| fig-cap: "With small samples, priors prevent overfitting"
# Demonstrate regularization benefit
set.seed(123)
# True DGP: AR(1)
T_obs <- 30
phi_true <- 0.7
y <- numeric(T_obs)
y[1] <- rnorm(1)
for (t in 2:T_obs) {
y[t] <- phi_true * y[t-1] + rnorm(1, 0, 0.5)
}
# Frequentist estimate
freq_est <- ar(y, order.max = 1, method = "ols")$ar
# Bayesian with different priors
# Prior 1: Tight prior at 0 (strong shrinkage)
# Prior 2: Moderate prior at 0.5
# Prior 3: Vague prior
# Simple conjugate normal prior on phi
bayesian_ar1 <- function(y, prior_mean, prior_var) {
T_obs <- length(y)
y_lag <- y[1:(T_obs-1)]
y_curr <- y[2:T_obs]
# OLS quantities
xtx <- sum(y_lag^2)
xty <- sum(y_lag * y_curr)
# Posterior (assuming known sigma = 1 for simplicity)
sigma2 <- var(y_curr - (xty/xtx) * y_lag)
post_var <- 1 / (1/prior_var + xtx/sigma2)
post_mean <- post_var * (prior_mean/prior_var + xty/sigma2)
c(mean = post_mean, sd = sqrt(post_var))
}
results <- data.frame(
method = c("Frequentist (OLS)", "Bayes: Tight N(0, 0.1)",
"Bayes: Moderate N(0.5, 0.5)", "Bayes: Vague N(0, 10)"),
estimate = c(
freq_est,
bayesian_ar1(y, 0, 0.1)[1],
bayesian_ar1(y, 0.5, 0.5)[1],
bayesian_ar1(y, 0, 10)[1]
)
)
ggplot(results, aes(x = method, y = estimate)) +
geom_point(size = 4, color = "#3498db") +
geom_hline(yintercept = phi_true, linetype = "dashed", color = "#e74c3c") +
annotate("text", x = 0.5, y = phi_true + 0.05,
label = paste0("True φ = ", phi_true), hjust = 0, color = "#e74c3c") +
coord_flip() +
labs(title = "Frequentist vs. Bayesian Estimates with Small Sample (T=30)",
subtitle = "Different priors lead to different shrinkage",
x = "", y = expression(hat(phi)))
```
## Challenge 2: Incorporating Prior Information
In macro, we often have strong theoretical priors:
- Interest rate semi-elasticity of money demand: around -0.05 to -0.15
- Labor share in production: around 0.6-0.7
- Intertemporal elasticity of substitution: around 0.5-2
**Bayesian solution**: Formally incorporate this information through priors.
## Challenge 3: Full Uncertainty Quantification
Frequentist confidence intervals answer: "If I repeated this procedure many times, 95% of intervals would contain the true value."
Bayesian credible intervals answer: "Given the data, there's a 95% probability the parameter lies in this interval."
The Bayesian interpretation is often what researchers actually want.
```{r}
#| label: uncertainty
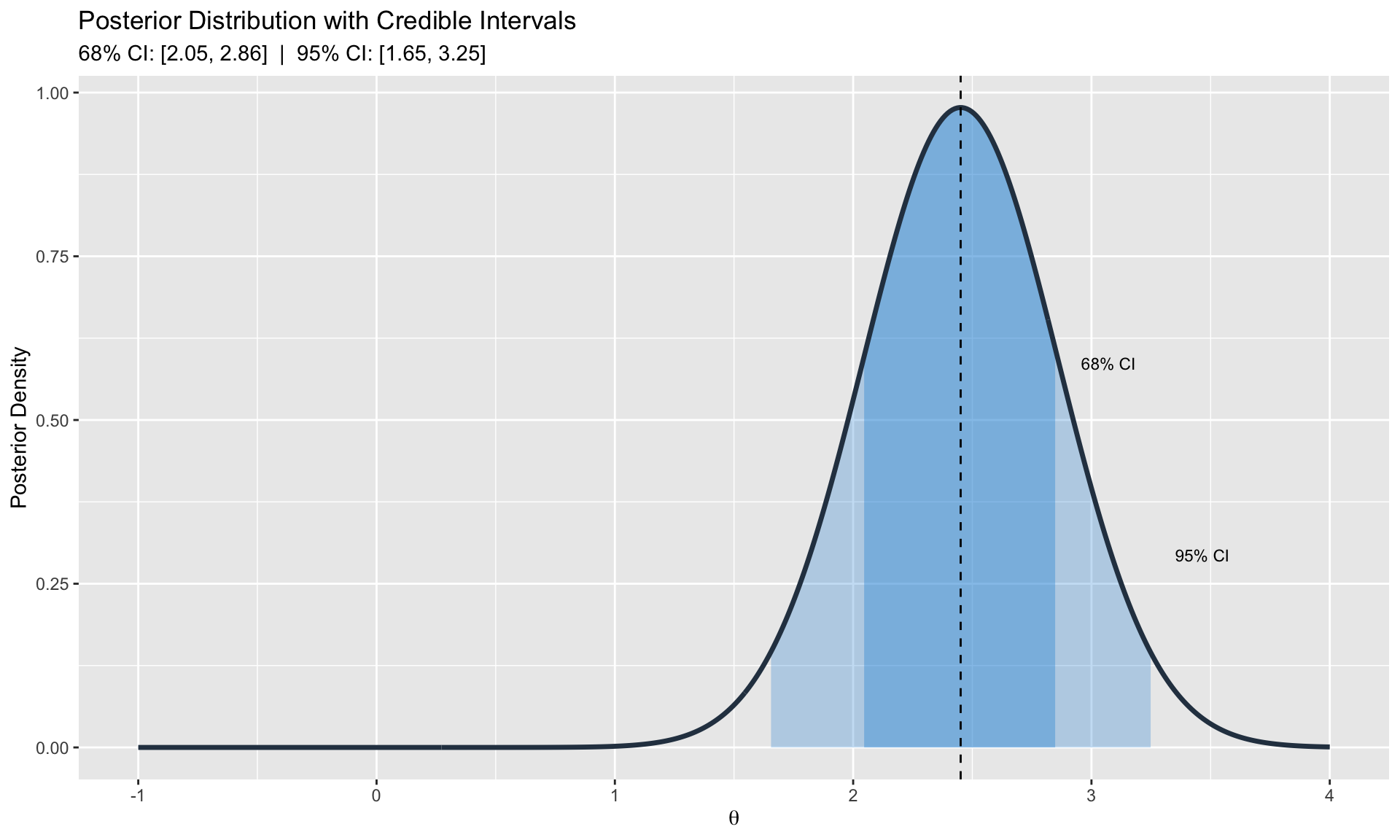
#| fig-cap: "Full posterior distributions quantify parameter uncertainty"
# Show posterior distribution with credible intervals
theta_grid <- seq(-1, 4, length.out = 500)
post_density <- dnorm(theta_grid, post_mean, sqrt(post_var))
# Credible intervals
ci_68 <- qnorm(c(0.16, 0.84), post_mean, sqrt(post_var))
ci_95 <- qnorm(c(0.025, 0.975), post_mean, sqrt(post_var))
ci_data <- data.frame(
x = theta_grid,
y = post_density,
in_68 = theta_grid >= ci_68[1] & theta_grid <= ci_68[2],
in_95 = theta_grid >= ci_95[1] & theta_grid <= ci_95[2]
)
ggplot(ci_data, aes(x = x, y = y)) +
geom_ribbon(data = filter(ci_data, in_95),
aes(ymin = 0, ymax = y), fill = "#3498db", alpha = 0.3) +
geom_ribbon(data = filter(ci_data, in_68),
aes(ymin = 0, ymax = y), fill = "#3498db", alpha = 0.4) +
geom_line(size = 1.2, color = "#2c3e50") +
geom_vline(xintercept = post_mean, linetype = "dashed") +
annotate("text", x = ci_95[2] + 0.1, y = max(post_density) * 0.3,
label = "95% CI", hjust = 0, size = 3) +
annotate("text", x = ci_68[2] + 0.1, y = max(post_density) * 0.6,
label = "68% CI", hjust = 0, size = 3) +
labs(title = "Posterior Distribution with Credible Intervals",
subtitle = paste0("68% CI: [", round(ci_68[1], 2), ", ", round(ci_68[2], 2),
"] | 95% CI: [", round(ci_95[1], 2), ", ", round(ci_95[2], 2), "]"),
x = expression(theta), y = "Posterior Density")
```
# Conjugate Priors {.unnumbered}
When the prior and posterior belong to the same family, we have **conjugate priors**. This allows analytical solutions without MCMC.
## Normal-Normal Conjugacy
**Model**: $Y_i | \mu \sim N(\mu, \sigma^2)$ with $\sigma^2$ known
**Prior**: $\mu \sim N(\mu_0, \tau_0^2)$
**Posterior**: $\mu | Y \sim N(\mu_n, \tau_n^2)$
where:
$$
\tau_n^2 = \left(\frac{1}{\tau_0^2} + \frac{n}{\sigma^2}\right)^{-1}, \quad
\mu_n = \tau_n^2 \left(\frac{\mu_0}{\tau_0^2} + \frac{n\bar{Y}}{\sigma^2}\right)
$$
**Interpretation**: Posterior mean is a precision-weighted average of prior mean and sample mean.
## Normal-Inverse-Gamma (Unknown Variance)
**Model**: $Y_i | \mu, \sigma^2 \sim N(\mu, \sigma^2)$
**Prior**:
$$
\mu | \sigma^2 \sim N(\mu_0, \sigma^2/\kappa_0), \quad
\sigma^2 \sim \text{Inv-Gamma}(\alpha_0, \beta_0)
$$
**Posterior**: Same family with updated parameters.
## Multivariate Normal-Inverse-Wishart
For VAR models with coefficient matrix $B$ and covariance $\Sigma$:
**Prior**:
$$
\text{vec}(B) | \Sigma \sim N(\text{vec}(B_0), \Sigma \otimes V_0), \quad
\Sigma \sim \text{Inv-Wishart}(S_0, \nu_0)
$$
**Posterior**: Same family—this is the foundation of Bayesian VARs.
```{r}
#| label: conjugate-table
# Summary table of conjugate priors
conjugate_table <- data.frame(
Likelihood = c("Normal (known var)", "Normal (unknown var)",
"Bernoulli/Binomial", "Poisson", "Multivariate Normal"),
Prior = c("Normal", "Normal-Inv-Gamma", "Beta", "Gamma", "Normal-Inv-Wishart"),
Posterior = c("Normal", "Normal-Inv-Gamma", "Beta", "Gamma", "Normal-Inv-Wishart"),
`Use Case` = c("Mean estimation", "Regression coefficients",
"Proportions", "Count data", "VAR coefficients")
)
knitr::kable(conjugate_table, caption = "Common Conjugate Prior Families")
```
# MCMC Methods {.unnumbered}
When posteriors don't have closed forms, we use **Markov Chain Monte Carlo (MCMC)** to draw samples from the posterior.
## The Core Idea
Instead of computing $p(\theta|Y)$ analytically, we:
1. Construct a Markov chain whose stationary distribution is $p(\theta|Y)$
2. Run the chain long enough to reach stationarity
3. Use the samples to approximate posterior quantities
$$
E[g(\theta)|Y] \approx \frac{1}{G} \sum_{g=1}^{G} g(\theta^{(g)})
$$
## Metropolis-Hastings Algorithm
The most general MCMC algorithm.
**Algorithm**:
1. Start at $\theta^{(0)}$
2. For $g = 1, \ldots, G$:
a. Propose $\theta^* \sim q(\theta^* | \theta^{(g-1)})$ (proposal distribution)
b. Compute acceptance probability:
$$
\alpha = \min\left(1, \frac{p(\theta^*|Y) \cdot q(\theta^{(g-1)}|\theta^*)}{p(\theta^{(g-1)}|Y) \cdot q(\theta^*|\theta^{(g-1)})}\right)
$$
c. Accept with probability $\alpha$: set $\theta^{(g)} = \theta^*$; otherwise $\theta^{(g)} = \theta^{(g-1)}$
```{r}
#| label: mh-algorithm
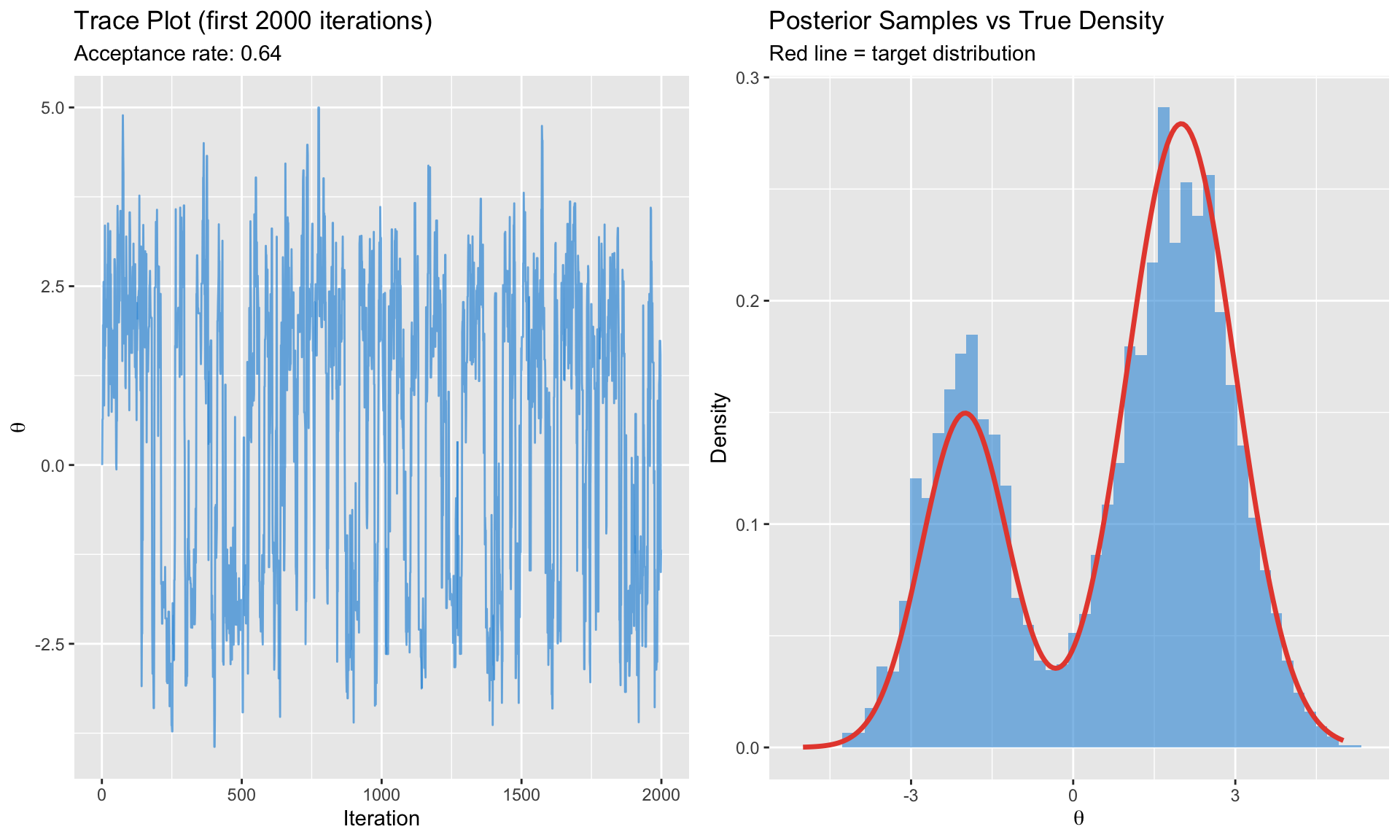
#| fig-cap: "Metropolis-Hastings sampling from a bimodal posterior"
# Demonstrate MH on a bimodal target
target_log <- function(x) {
# Mixture of two normals
log(0.3 * dnorm(x, -2, 0.8) + 0.7 * dnorm(x, 2, 1))
}
metropolis_hastings <- function(n_iter, start, proposal_sd) {
samples <- numeric(n_iter)
samples[1] <- start
accepted <- 0
for (i in 2:n_iter) {
# Propose
proposal <- rnorm(1, samples[i-1], proposal_sd)
# Acceptance probability (log scale)
log_alpha <- target_log(proposal) - target_log(samples[i-1])
if (log(runif(1)) < log_alpha) {
samples[i] <- proposal
accepted <- accepted + 1
} else {
samples[i] <- samples[i-1]
}
}
list(samples = samples, acceptance_rate = accepted / (n_iter - 1))
}
# Run MH
n_iter <- 10000
mh_result <- metropolis_hastings(n_iter, start = 0, proposal_sd = 1.5)
# Plot
par(mfrow = c(1, 2))
# Trace plot
trace_df <- data.frame(
iteration = 1:n_iter,
value = mh_result$samples
)
p1 <- ggplot(trace_df[1:2000, ], aes(x = iteration, y = value)) +
geom_line(alpha = 0.7, color = "#3498db") +
labs(title = "Trace Plot (first 2000 iterations)",
subtitle = paste0("Acceptance rate: ", round(mh_result$acceptance_rate, 2)),
x = "Iteration", y = expression(theta))
# Histogram vs true density
x_grid <- seq(-5, 5, length.out = 200)
true_density <- 0.3 * dnorm(x_grid, -2, 0.8) + 0.7 * dnorm(x_grid, 2, 1)
p2 <- ggplot() +
geom_histogram(data = data.frame(x = mh_result$samples[1001:n_iter]),
aes(x = x, y = after_stat(density)),
bins = 50, fill = "#3498db", alpha = 0.6) +
geom_line(data = data.frame(x = x_grid, y = true_density),
aes(x = x, y = y), color = "#e74c3c", size = 1.2) +
labs(title = "Posterior Samples vs True Density",
subtitle = "Red line = target distribution",
x = expression(theta), y = "Density")
gridExtra::grid.arrange(p1, p2, ncol = 2)
```
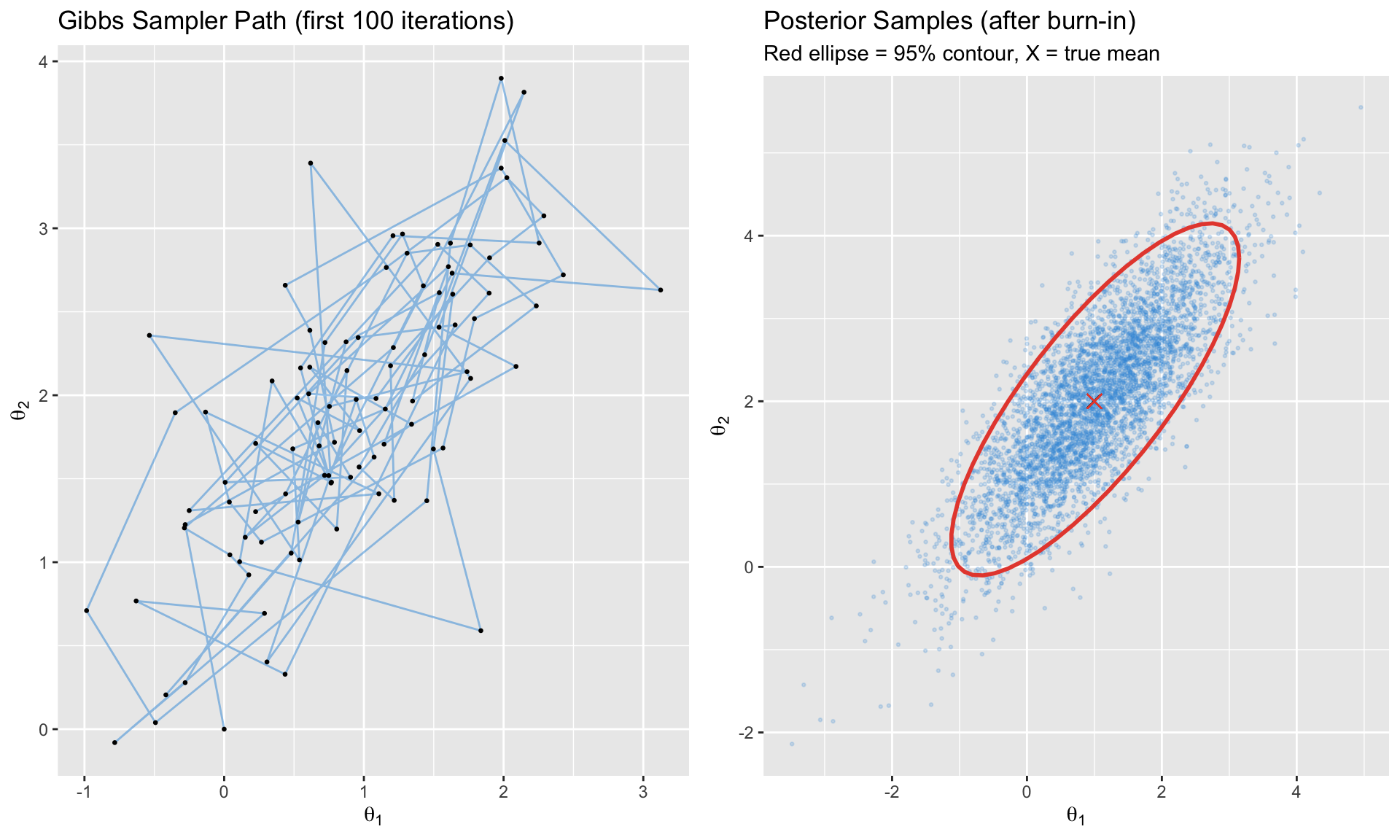
## Gibbs Sampling
A special case of MH where we sample each parameter **conditionally** on the others.
**Algorithm** (for $\theta = (\theta_1, \theta_2)$):
1. Start at $(\theta_1^{(0)}, \theta_2^{(0)})$
2. For $g = 1, \ldots, G$:
a. Draw $\theta_1^{(g)} \sim p(\theta_1 | \theta_2^{(g-1)}, Y)$
b. Draw $\theta_2^{(g)} \sim p(\theta_2 | \theta_1^{(g)}, Y)$
**Advantage**: No tuning required (acceptance rate = 100%)
**Requirement**: Must know conditional distributions
```{r}
#| label: gibbs-sampling
#| fig-cap: "Gibbs sampling from a bivariate normal"
# Gibbs sampler for bivariate normal
gibbs_bivariate_normal <- function(n_iter, mu, Sigma) {
# Target: N(mu, Sigma)
rho <- Sigma[1,2] / sqrt(Sigma[1,1] * Sigma[2,2])
sd1 <- sqrt(Sigma[1,1])
sd2 <- sqrt(Sigma[2,2])
samples <- matrix(0, n_iter, 2)
samples[1, ] <- c(0, 0) # start
for (i in 2:n_iter) {
# Draw x1 | x2
cond_mean1 <- mu[1] + rho * sd1/sd2 * (samples[i-1, 2] - mu[2])
cond_sd1 <- sd1 * sqrt(1 - rho^2)
samples[i, 1] <- rnorm(1, cond_mean1, cond_sd1)
# Draw x2 | x1
cond_mean2 <- mu[2] + rho * sd2/sd1 * (samples[i, 1] - mu[1])
cond_sd2 <- sd2 * sqrt(1 - rho^2)
samples[i, 2] <- rnorm(1, cond_mean2, cond_sd2)
}
samples
}
# Run Gibbs
mu <- c(1, 2)
Sigma <- matrix(c(1, 0.8, 0.8, 1), 2, 2)
gibbs_samples <- gibbs_bivariate_normal(5000, mu, Sigma)
# Plot trajectory and samples
gibbs_df <- data.frame(
x1 = gibbs_samples[, 1],
x2 = gibbs_samples[, 2],
iteration = 1:nrow(gibbs_samples)
)
# First 100 iterations showing path
p1 <- ggplot(gibbs_df[1:100, ], aes(x = x1, y = x2)) +
geom_path(alpha = 0.5, color = "#3498db") +
geom_point(size = 0.5) +
labs(title = "Gibbs Sampler Path (first 100 iterations)",
x = expression(theta[1]), y = expression(theta[2]))
# All samples
p2 <- ggplot(gibbs_df[501:5000, ], aes(x = x1, y = x2)) +
geom_point(alpha = 0.2, size = 0.5, color = "#3498db") +
stat_ellipse(level = 0.95, color = "#e74c3c", size = 1) +
geom_point(aes(x = mu[1], y = mu[2]), color = "#e74c3c", size = 3, shape = 4) +
labs(title = "Posterior Samples (after burn-in)",
subtitle = "Red ellipse = 95% contour, X = true mean",
x = expression(theta[1]), y = expression(theta[2]))
gridExtra::grid.arrange(p1, p2, ncol = 2)
```
## Gibbs for Bayesian Regression
For the normal linear model $Y = X\beta + \varepsilon$ with $\varepsilon \sim N(0, \sigma^2 I)$:
**Block 1**: Draw $\beta | \sigma^2, Y$
$$
\beta | \sigma^2, Y \sim N\left(V_n X'Y, \sigma^2 V_n\right), \quad V_n = (V_0^{-1} + X'X)^{-1}
$$
**Block 2**: Draw $\sigma^2 | \beta, Y$
$$
\sigma^2 | \beta, Y \sim \text{Inv-Gamma}\left(\frac{\nu_0 + n}{2}, \frac{\nu_0 s_0^2 + (Y-X\beta)'(Y-X\beta)}{2}\right)
$$
## Hamiltonian Monte Carlo (HMC)
Modern MCMC that uses gradient information for efficient exploration.
**Key idea**: Treat the parameter space as a physical system. The negative log-posterior is "potential energy." Add "momentum" variables and simulate Hamiltonian dynamics.
**Advantages**:
- Much more efficient for high-dimensional problems
- Fewer iterations needed for convergence
- Better exploration of complex posteriors
**Implementation**: Use Stan (via `rstan` or `cmdstanr`) or PyMC
```{r}
#| label: mcmc-comparison
# Comparison table of MCMC methods
mcmc_table <- data.frame(
Method = c("Metropolis-Hastings", "Gibbs Sampling", "HMC/NUTS"),
Tuning = c("Proposal variance", "None (if conjugate)", "Step size, path length"),
Efficiency = c("Low-Medium", "Medium", "High"),
`Best For` = c("Simple models, low dimensions",
"Conjugate models, block structure",
"High dimensions, complex posteriors"),
Software = c("Manual, MCMCpack", "Manual, JAGS", "Stan, PyMC")
)
knitr::kable(mcmc_table, caption = "Comparison of MCMC Methods")
```
# Convergence Diagnostics {.unnumbered}
MCMC is only useful if the chain has converged to the posterior. Always check!
## Visual Diagnostics
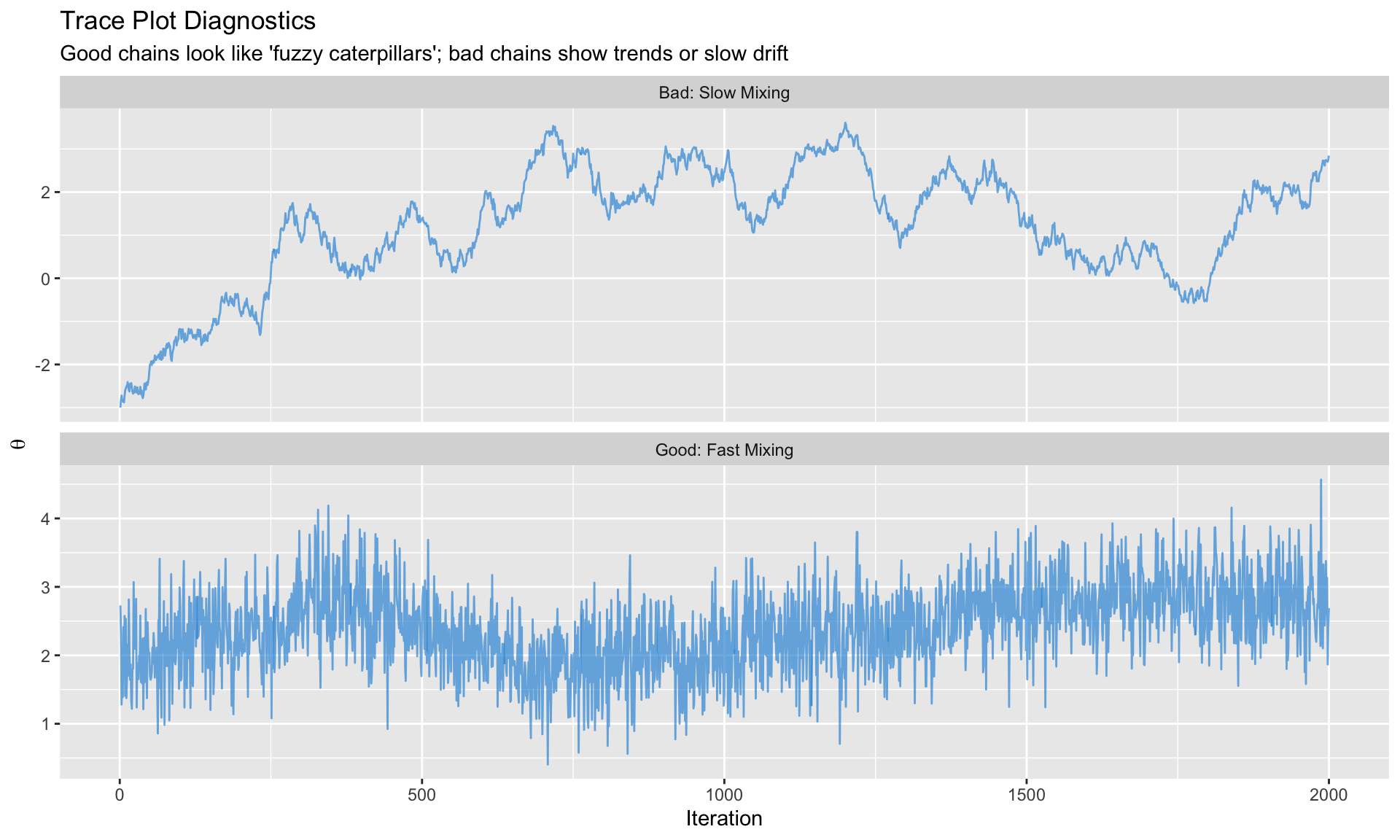
### Trace Plots
The chain should look like a "fuzzy caterpillar"—no trends, quick mixing.
```{r}
#| label: trace-plots
#| fig-cap: "Good vs. bad trace plots"
# Good chain
good_chain <- cumsum(rnorm(2000, 0, 1)) * 0.02 + rnorm(2000, 2, 0.5)
# Bad chain (slow mixing)
bad_chain <- numeric(2000)
bad_chain[1] <- -3
for (i in 2:2000) {
bad_chain[i] <- 0.995 * bad_chain[i-1] + rnorm(1, 0.01, 0.1)
}
trace_comparison <- data.frame(
iteration = rep(1:2000, 2),
value = c(good_chain, bad_chain),
chain = rep(c("Good: Fast Mixing", "Bad: Slow Mixing"), each = 2000)
)
ggplot(trace_comparison, aes(x = iteration, y = value)) +
geom_line(alpha = 0.7, color = "#3498db") +
facet_wrap(~chain, scales = "free_y", ncol = 1) +
labs(title = "Trace Plot Diagnostics",
subtitle = "Good chains look like 'fuzzy caterpillars'; bad chains show trends or slow drift",
x = "Iteration", y = expression(theta))
```
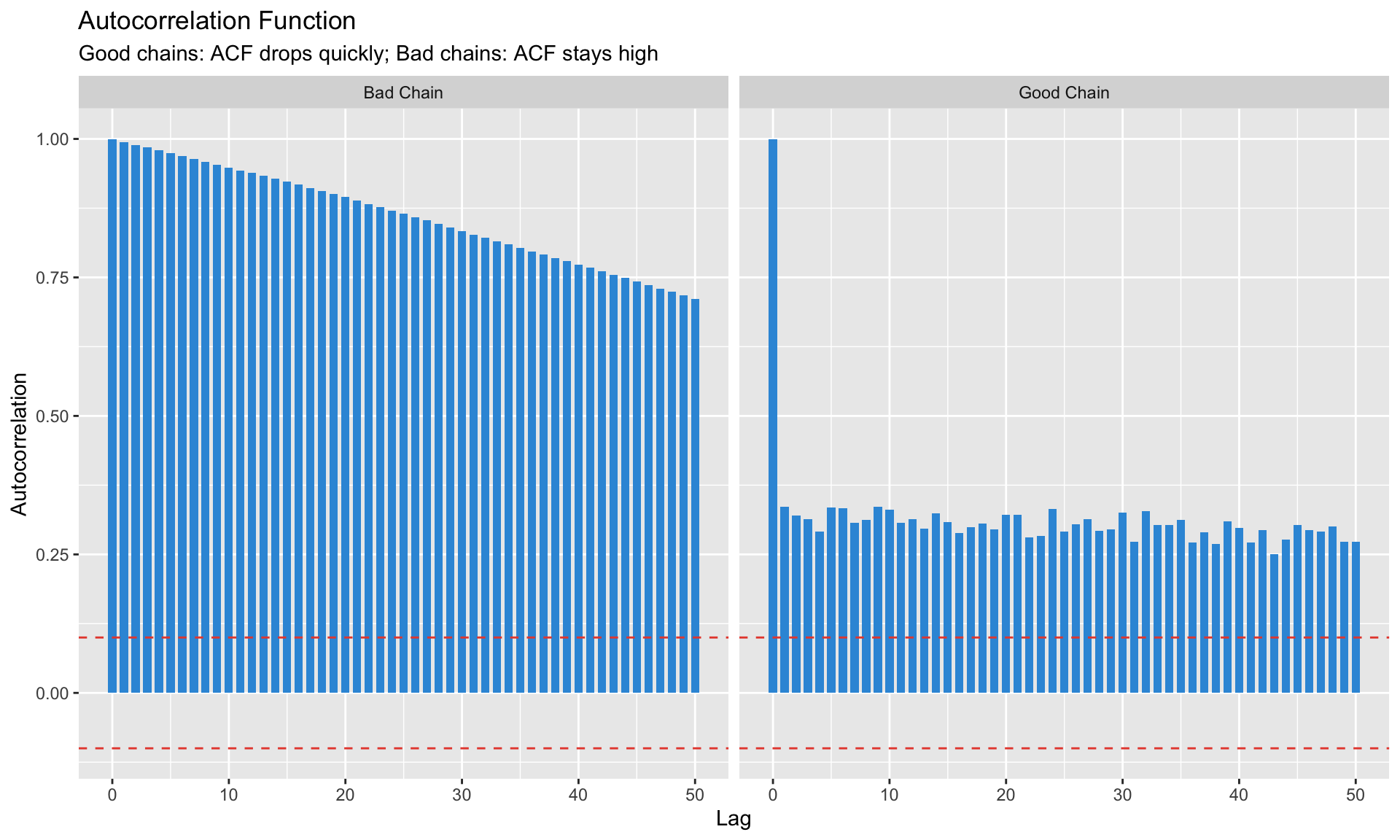
### Autocorrelation Plots
Samples should be approximately independent. High autocorrelation = inefficient sampling.
```{r}
#| label: acf-plots
#| fig-cap: "Autocorrelation should decay quickly"
# Calculate ACF
acf_good <- acf(good_chain, lag.max = 50, plot = FALSE)
acf_bad <- acf(bad_chain, lag.max = 50, plot = FALSE)
acf_df <- data.frame(
lag = rep(0:50, 2),
acf = c(acf_good$acf, acf_bad$acf),
chain = rep(c("Good Chain", "Bad Chain"), each = 51)
)
ggplot(acf_df, aes(x = lag, y = acf)) +
geom_col(fill = "#3498db", width = 0.7) +
geom_hline(yintercept = c(-0.1, 0.1), linetype = "dashed", color = "#e74c3c") +
facet_wrap(~chain, ncol = 2) +
labs(title = "Autocorrelation Function",
subtitle = "Good chains: ACF drops quickly; Bad chains: ACF stays high",
x = "Lag", y = "Autocorrelation")
```
## Quantitative Diagnostics
### Gelman-Rubin $\hat{R}$ Statistic
Run **multiple chains** from different starting points. Compare within-chain variance to between-chain variance.
$$
\hat{R} = \sqrt{\frac{\text{Var}(\theta | Y)_{\text{estimate}}}{\text{Within-chain variance}}}
$$
**Rule**: $\hat{R} < 1.1$ (ideally $< 1.05$)
### Effective Sample Size (ESS)
Accounts for autocorrelation. With $G$ draws and autocorrelation $\rho_k$ at lag $k$:
$$
\text{ESS} = \frac{G}{1 + 2\sum_{k=1}^{\infty} \rho_k}
$$
**Rule**: ESS $> 400$ for reliable posterior summaries
```{r}
#| label: ess-calculation
# Calculate ESS (simplified)
calculate_ess <- function(chain) {
n <- length(chain)
acf_vals <- acf(chain, lag.max = min(n-1, 100), plot = FALSE)$acf[-1]
# Sum until first negative autocorrelation
sum_rho <- 0
for (rho in acf_vals) {
if (rho < 0) break
sum_rho <- sum_rho + rho
}
n / (1 + 2 * sum_rho)
}
cat("Good chain ESS:", round(calculate_ess(good_chain)), "out of", length(good_chain), "\n")
cat("Bad chain ESS:", round(calculate_ess(bad_chain)), "out of", length(bad_chain), "\n")
```
### Diagnostic Summary Table
```{r}
#| label: diagnostic-table
diagnostic_table <- data.frame(
Diagnostic = c("Trace plot", "R-hat", "ESS", "Geweke", "Autocorrelation"),
Target = c("No trends, good mixing", "< 1.1 (ideally < 1.05)",
"> 400", "p > 0.05", "Quick decay"),
`What It Catches` = c("Non-stationarity", "Between-chain disagreement",
"High autocorrelation", "Drift in mean",
"Inefficient sampling")
)
knitr::kable(diagnostic_table, caption = "MCMC Convergence Diagnostics")
```
# Model Comparison {.unnumbered}
Bayesian model comparison answers: "Which model is better supported by the data?"
## Marginal Likelihood
The probability of the data under the model:
$$
p(Y | M) = \int p(Y | \theta, M) p(\theta | M) d\theta
$$
This is the denominator in Bayes' theorem—it averages the likelihood over the prior.
## Bayes Factors
Compare two models:
$$
BF_{12} = \frac{p(Y | M_1)}{p(Y | M_2)}
$$
| log(BF) | Interpretation |
|---------|----------------|
| 0-1 | Not worth mentioning |
| 1-3 | Positive evidence |
| 3-5 | Strong evidence |
| > 5 | Very strong evidence |
## Computing Marginal Likelihood
**Challenge**: The integral is often intractable.
**Methods**:
1. **Analytical** (for conjugate models like NIW-BVAR)
2. **Harmonic Mean** (easy but unreliable—avoid!)
3. **Chib's Method** (for Gibbs samplers)
4. **Bridge Sampling** (general, reliable)
```{r}
#| label: model-comparison
#| fig-cap: "Model comparison via marginal likelihood"
# Simple example: comparing models with different priors
# Model 1: Tight prior (N(0, 0.5))
# Model 2: Vague prior (N(0, 5))
# For normal-normal model, marginal likelihood is available analytically
# p(Y | prior) = N(Y | prior_mean, prior_var + data_var/n)
y_obs <- c(1.2, 1.5, 0.8, 1.1, 1.4) # sample data
n <- length(y_obs)
y_bar <- mean(y_obs)
sigma2 <- 1 # known variance
# Marginal likelihood under each prior
log_ml <- function(prior_mean, prior_var, y_bar, n, sigma2) {
marginal_var <- prior_var + sigma2/n
dnorm(y_bar, prior_mean, sqrt(marginal_var), log = TRUE)
}
models <- data.frame(
model = c("M1: Tight prior N(0, 0.5)", "M2: Vague prior N(0, 5)",
"M3: Centered prior N(1, 1)"),
prior_mean = c(0, 0, 1),
prior_var = c(0.5, 5, 1)
)
models$log_ml <- mapply(log_ml, models$prior_mean, models$prior_var,
MoreArgs = list(y_bar = y_bar, n = n, sigma2 = sigma2))
models$posterior_prob <- exp(models$log_ml) / sum(exp(models$log_ml))
# Bayes factors relative to M1
models$BF_vs_M1 <- exp(models$log_ml - models$log_ml[1])
cat("Data: y_bar =", round(y_bar, 2), "\n\n")
print(models[, c("model", "log_ml", "BF_vs_M1", "posterior_prob")])
```
## Information Criteria
**DIC (Deviance Information Criterion)**:
$$
DIC = \bar{D}(\theta) + p_D
$$
where $\bar{D}$ is average deviance and $p_D$ is effective number of parameters.
**WAIC (Widely Applicable IC)**:
- Fully Bayesian
- Asymptotically equivalent to leave-one-out cross-validation
- More reliable than DIC
**Rule**: Lower is better (like AIC/BIC)
# Practical Implementation {.unnumbered}
## R: Using coda for Diagnostics
```{r}
#| label: coda-example
#| eval: false
library(coda)
# Convert MCMC output to coda object
mcmc_chain <- as.mcmc(posterior_draws)
# Diagnostics
plot(mcmc_chain) # Trace and density
autocorr.plot(mcmc_chain) # ACF
effectiveSize(mcmc_chain) # ESS
geweke.diag(mcmc_chain) # Geweke test
# For multiple chains
mcmc_list <- mcmc.list(chain1, chain2, chain3)
gelman.diag(mcmc_list) # R-hat
gelman.plot(mcmc_list) # R-hat over iterations
```
## R: Using BVAR Package
```{r}
#| label: bvar-example
#| eval: false
library(BVAR)
# Estimate Bayesian VAR with Minnesota prior
bvar_model <- bvar(
data = macro_data,
lags = 4,
n_draw = 20000,
n_burn = 5000,
priors = bv_minnesota(
lambda = bv_lambda(mode = 0.2, sd = 0.4, min = 0.0001, max = 5),
alpha = bv_alpha(mode = 2)
)
)
# Check convergence
plot(bvar_model) # trace plots
# Posterior summaries
summary(bvar_model)
# IRFs with credible bands
irf_results <- irf(bvar_model, horizon = 20)
plot(irf_results)
```
## Python: Using PyMC
```python
import pymc as pm
import numpy as np
import arviz as az
# Example: Bayesian linear regression
with pm.Model() as model:
# Priors
alpha = pm.Normal("alpha", mu=0, sigma=10)
beta = pm.Normal("beta", mu=0, sigma=10)
sigma = pm.HalfNormal("sigma", sigma=1)
# Likelihood
mu = alpha + beta * X
y_obs = pm.Normal("y", mu=mu, sigma=sigma, observed=y)
# Sample
trace = pm.sample(2000, tune=1000, cores=2)
# Diagnostics
az.plot_trace(trace)
az.summary(trace)
az.plot_posterior(trace)
# Model comparison
az.waic(trace)
az.loo(trace)
```
# Summary {.unnumbered}
Key takeaways from this module:
1. **Bayesian inference** combines prior beliefs with data through Bayes' theorem: Posterior ∝ Likelihood × Prior
2. **Why Bayesian for macro**:
- Small samples → priors prevent overfitting
- Prior information → formally incorporated
- Full uncertainty quantification → credible intervals
3. **Conjugate priors** allow analytical posteriors (Normal-Inverse-Wishart for VARs)
4. **MCMC methods** sample from posteriors when analytical solutions don't exist:
- Metropolis-Hastings: general purpose
- Gibbs sampling: for conditional sampling
- HMC/NUTS: efficient for high dimensions (Stan, PyMC)
5. **Always check convergence**:
- Trace plots (fuzzy caterpillar)
- R-hat < 1.1
- ESS > 400
6. **Model comparison** via marginal likelihood, Bayes factors, or information criteria (DIC, WAIC)
---
## Key References
**Foundational**:
- Koop (2003). *Bayesian Econometrics*
- Greenberg (2008). *Introduction to Bayesian Econometrics*
- Gelman et al. (2013). *Bayesian Data Analysis* (3rd ed.)
**MCMC Methods**:
- Chib & Greenberg (1995). "Understanding the Metropolis-Hastings Algorithm." *The American Statistician*
- Gelfand & Smith (1990). "Sampling-Based Approaches to Calculating Marginal Densities." *JASA*
**Diagnostics**:
- Gelman & Rubin (1992). "Inference from Iterative Simulation Using Multiple Sequences." *Statistical Science*
- Vehtari et al. (2021). "Rank-Normalization, Folding, and Localization: An Improved R-hat." *Bayesian Analysis*
**Model Comparison**:
- Kass & Raftery (1995). "Bayes Factors." *JASA*
- Chib (1995). "Marginal Likelihood from the Gibbs Output." *JASA*
**Software**:
- Stan Development Team. [Stan User's Guide](https://mc-stan.org/users/documentation/)
- PyMC Developers. [PyMC Documentation](https://www.pymc.io/)
---
*Next: [Module 8: Bayesian VARs](08_bayesian_var.qmd)*