---
title: "Bayesian Panel Methods"
subtitle: "Hierarchical Models, Panel VARs, and Cross-Sectional Dependence"
---
```{r}
#| label: setup
#| include: false
library(tidyverse)
library(mvtnorm)
set.seed(42)
```
# The Panel Data Challenge {.unnumbered}
Cross-country macroeconomic panels present a fundamental tension:
**Heterogeneity**: Countries differ in institutions, structures, and responses
**Limited Data**: Each country has finite time series (T = 84 quarters)
**Efficiency**: Ignoring cross-country information wastes data
::: {.callout-note}
## The Core Question
How do we **pool information** across countries while **respecting heterogeneity**?
:::
## Three Approaches
| Approach | Assumption | Problem |
|----------|------------|---------|
| **Pooled** | All countries identical | Ignores heterogeneity |
| **Country-by-country** | Countries unrelated | Inefficient, noisy |
| **Hierarchical Bayes** | Countries related but different | Best of both worlds |
With N = 46 countries and T = 84 quarters, hierarchical Bayesian methods offer the natural solution.
# Hierarchical Models: The Framework {.unnumbered}
## The Multilevel Structure
Consider a simple panel regression:
$$
y_{it} = x_{it}'\beta_i + \varepsilon_{it}, \quad \varepsilon_{it} \sim N(0, \sigma^2_i)
$$
**Level 1 (Within-Country)**:
$$
\beta_i | \mu, \Omega \sim N(\mu, \Omega)
$$
**Level 2 (Across-Countries)**:
$$
\mu \sim N(\mu_0, V_0), \quad \Omega \sim \text{Inverse-Wishart}(\Omega_0, \nu_0)
$$
The country-specific coefficients $\beta_i$ are **drawn from a common distribution** with unknown mean $\mu$ and variance $\Omega$.
## Shrinkage and Partial Pooling
The posterior for $\beta_i$ combines:
1. **Country-specific data** (likelihood)
2. **Cross-country information** (hierarchical prior)
$$
\hat{\beta}_i^{\text{Bayes}} = \lambda_i \hat{\beta}_i^{\text{OLS}} + (1 - \lambda_i) \bar{\beta}
$$
where $\lambda_i \in [0,1]$ depends on:
- **Precision of country data**: More data → less shrinkage
- **Cross-country heterogeneity**: More variance in $\Omega$ → less shrinkage
```{r}
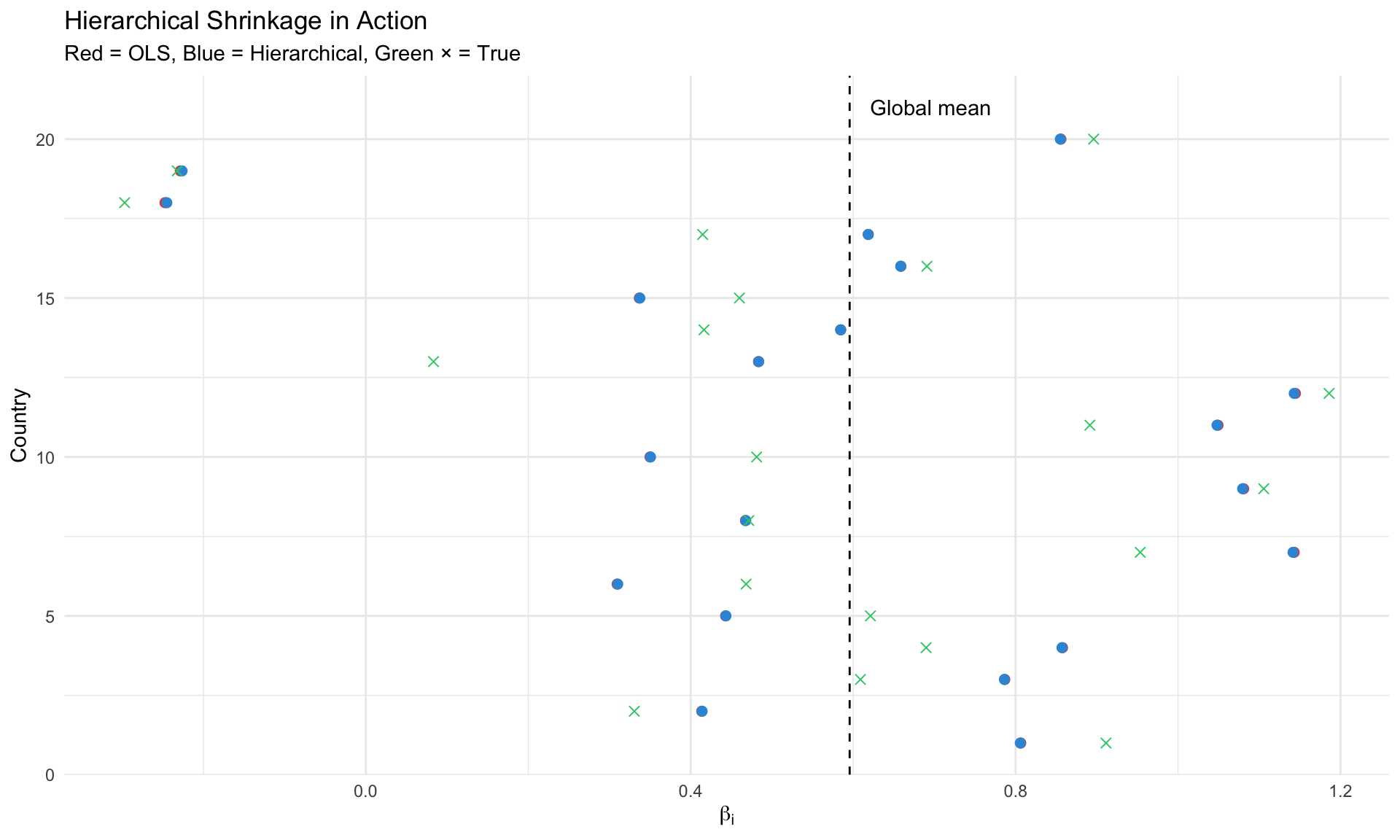
#| label: shrinkage-illustration
#| fig-cap: "Hierarchical shrinkage: country estimates pulled toward global mean"
# Simulate hierarchical data
N <- 20 # countries
T_obs <- 50 # periods per country
# True hierarchical structure
mu_true <- 0.5
omega_true <- 0.3^2
sigma_true <- 1
# Country-specific coefficients
beta_true <- rnorm(N, mu_true, sqrt(omega_true))
# Generate data
data_hier <- map_dfr(1:N, function(i) {
x <- rnorm(T_obs)
y <- beta_true[i] * x + rnorm(T_obs, 0, sigma_true)
tibble(country = i, x = x, y = y)
})
# OLS estimates by country
ols_estimates <- data_hier %>%
group_by(country) %>%
summarise(
beta_ols = coef(lm(y ~ x - 1))[1],
se_ols = summary(lm(y ~ x - 1))$coefficients[1, 2],
n = n()
)
# Hierarchical shrinkage (empirical Bayes approximation)
grand_mean <- mean(ols_estimates$beta_ols)
between_var <- var(ols_estimates$beta_ols)
within_var <- mean(ols_estimates$se_ols^2)
shrinkage_factor <- between_var / (between_var + within_var / T_obs)
ols_estimates <- ols_estimates %>%
mutate(
beta_hier = shrinkage_factor * beta_ols + (1 - shrinkage_factor) * grand_mean,
beta_true = beta_true
)
# Plot
ggplot(ols_estimates) +
geom_segment(aes(x = beta_ols, xend = beta_hier, y = country, yend = country),
arrow = arrow(length = unit(0.1, "cm")), color = "gray50") +
geom_point(aes(x = beta_ols, y = country), color = "#e74c3c", size = 2) +
geom_point(aes(x = beta_hier, y = country), color = "#3498db", size = 2) +
geom_point(aes(x = beta_true, y = country), shape = 4, size = 2, color = "#2ecc71") +
geom_vline(xintercept = grand_mean, linetype = "dashed") +
annotate("text", x = grand_mean + 0.1, y = N + 1, label = "Global mean") +
labs(title = "Hierarchical Shrinkage in Action",
subtitle = "Red = OLS, Blue = Hierarchical, Green × = True",
x = expression(beta[i]), y = "Country") +
theme_minimal()
```
## Why Shrinkage Helps
**Key Insight**: Countries with less precise estimates get shrunk more toward the global mean.
| Country Type | Data Quality | Shrinkage | Result |
|--------------|--------------|-----------|--------|
| Data-rich (US, Germany) | High | Low | Near OLS |
| Data-poor (small EMs) | Low | High | Near global mean |
This is **not bias** — it's optimal use of information under the hierarchical structure.
# Gibbs Sampler for Hierarchical Panel {.unnumbered}
## The Algorithm
For the hierarchical linear model:
**Block 1**: Draw $\beta_i | y_i, \mu, \Omega, \sigma^2_i$ for each country
$$
\beta_i | \cdot \sim N\left( V_i^{-1}(X_i'y_i/\sigma^2_i + \Omega^{-1}\mu), V_i^{-1} \right)
$$
where $V_i = X_i'X_i/\sigma^2_i + \Omega^{-1}$
**Block 2**: Draw $\sigma^2_i | y_i, \beta_i$
$$
\sigma^2_i | \cdot \sim \text{Inverse-Gamma}\left( \frac{T_i}{2}, \frac{(y_i - X_i\beta_i)'(y_i - X_i\beta_i)}{2} \right)
$$
**Block 3**: Draw $\mu | \beta_1, \ldots, \beta_N, \Omega$
$$
\mu | \cdot \sim N\left( (N\Omega^{-1} + V_0^{-1})^{-1}(N\Omega^{-1}\bar{\beta} + V_0^{-1}\mu_0), (N\Omega^{-1} + V_0^{-1})^{-1} \right)
$$
**Block 4**: Draw $\Omega | \beta_1, \ldots, \beta_N, \mu$
$$
\Omega | \cdot \sim \text{Inverse-Wishart}\left( \Omega_0 + \sum_{i=1}^N (\beta_i - \mu)(\beta_i - \mu)', \nu_0 + N \right)
$$
```{r}
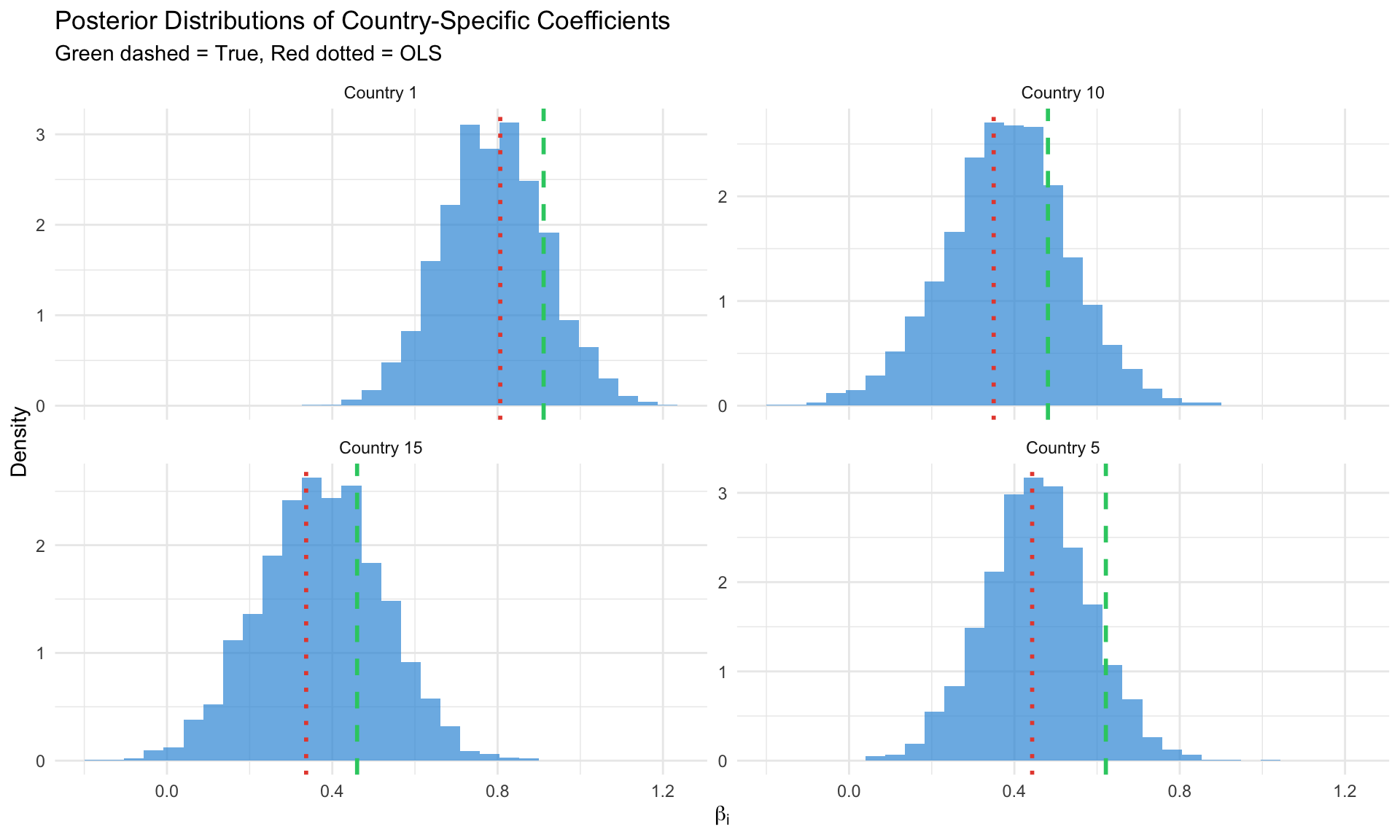
#| label: hierarchical-gibbs
#| fig-cap: "Posterior distributions from hierarchical panel model"
# Hierarchical Gibbs sampler for panel regression
hierarchical_panel_gibbs <- function(data, n_draw = 5000, n_burn = 1000) {
countries <- unique(data$country)
N <- length(countries)
K <- 1 # number of regressors (simplified)
# Organize data by country
country_data <- map(countries, function(c) {
d <- filter(data, country == c)
list(y = d$y, X = matrix(d$x, ncol = 1), T_i = nrow(d))
})
# Priors
mu0 <- 0
V0 <- 10
omega0 <- 1
nu0 <- 3
a0 <- 0.01; b0 <- 0.01 # for sigma^2
# Storage
beta_draws <- array(0, dim = c(N, K, n_draw))
mu_draws <- matrix(0, n_draw, K)
omega_draws <- numeric(n_draw)
sigma2_draws <- matrix(0, n_draw, N)
# Initialize
beta_curr <- rep(0, N)
sigma2_curr <- rep(1, N)
mu_curr <- 0
omega_curr <- 1
for (g in 1:(n_draw + n_burn)) {
# Block 1: Draw beta_i | rest
for (i in 1:N) {
X_i <- country_data[[i]]$X
y_i <- country_data[[i]]$y
T_i <- country_data[[i]]$T_i
V_i <- as.numeric(crossprod(X_i) / sigma2_curr[i] + 1/omega_curr)
m_i <- (crossprod(X_i, y_i) / sigma2_curr[i] + mu_curr/omega_curr) / V_i
beta_curr[i] <- rnorm(1, m_i, sqrt(1/V_i))
}
# Block 2: Draw sigma2_i | rest
for (i in 1:N) {
X_i <- country_data[[i]]$X
y_i <- country_data[[i]]$y
T_i <- country_data[[i]]$T_i
resid <- y_i - X_i * beta_curr[i]
a_post <- a0 + T_i/2
b_post <- b0 + sum(resid^2)/2
sigma2_curr[i] <- 1/rgamma(1, a_post, b_post)
}
# Block 3: Draw mu | rest
V_mu <- 1 / (N/omega_curr + 1/V0)
m_mu <- V_mu * (sum(beta_curr)/omega_curr + mu0/V0)
mu_curr <- rnorm(1, m_mu, sqrt(V_mu))
# Block 4: Draw omega | rest
nu_post <- nu0 + N
S_post <- omega0 + sum((beta_curr - mu_curr)^2)
omega_curr <- 1/rgamma(1, nu_post/2, S_post/2)
# Store
if (g > n_burn) {
idx <- g - n_burn
beta_draws[, , idx] <- beta_curr
mu_draws[idx, ] <- mu_curr
omega_draws[idx] <- omega_curr
sigma2_draws[idx, ] <- sigma2_curr
}
}
list(beta = beta_draws, mu = mu_draws, omega = omega_draws, sigma2 = sigma2_draws)
}
# Run on simulated data
hier_result <- hierarchical_panel_gibbs(data_hier, n_draw = 3000, n_burn = 1000)
# Posterior summary
beta_post_mean <- apply(hier_result$beta, 1, mean)
beta_post_sd <- apply(hier_result$beta, 1, sd)
# Compare to truth
comparison <- tibble(
country = 1:N,
true = beta_true,
ols = ols_estimates$beta_ols,
bayes_mean = beta_post_mean,
bayes_sd = beta_post_sd
)
# Plot posterior distributions for selected countries
selected <- c(1, 5, 10, 15)
post_df <- map_dfr(selected, function(i) {
tibble(
country = paste("Country", i),
value = hier_result$beta[i, 1, ],
true = beta_true[i],
ols = ols_estimates$beta_ols[i]
)
})
ggplot(post_df, aes(x = value)) +
geom_histogram(aes(y = after_stat(density)), bins = 30, fill = "#3498db", alpha = 0.7) +
geom_vline(aes(xintercept = true), color = "#2ecc71", linetype = "dashed", linewidth = 1) +
geom_vline(aes(xintercept = ols), color = "#e74c3c", linetype = "dotted", linewidth = 1) +
facet_wrap(~country, scales = "free_y") +
labs(title = "Posterior Distributions of Country-Specific Coefficients",
subtitle = "Green dashed = True, Red dotted = OLS",
x = expression(beta[i]), y = "Density") +
theme_minimal()
```
# Bayesian Panel VAR {.unnumbered}
## The Model Structure
For N countries, each with a K-variable VAR(p):
$$
y_{it} = c_i + A_{i,1} y_{i,t-1} + \cdots + A_{i,p} y_{i,t-p} + u_{it}
$$
**Hierarchical Prior**:
$$
\text{vec}(B_i) | \mu_B, \Omega_B \sim N(\mu_B, \Omega_B)
$$
where $B_i = [c_i, A_{i,1}, \ldots, A_{i,p}]'$ stacks all VAR coefficients for country $i$.
## Why Panel VAR?
| Feature | Separate VARs | Pooled VAR | Panel VAR |
|---------|---------------|------------|-----------|
| Heterogeneity | Yes | No | Yes |
| Sample size | T per country | N×T | N×T (shared info) |
| Curse of dimensionality | Severe | Moderate | Mitigated |
| Interpretation | Country-specific | Global average | Both |
With K = 3 variables, p = 2 lags, and T = 84:
- **Parameters per country**: 3 × (1 + 3×2) = 21
- **Observations per country**: 84 - 2 = 82
- **Degrees of freedom**: Tight!
Hierarchical pooling provides **effective sample size boost** without forcing homogeneity.
## Mean-Group Prior Structure
Following Canova & Ciccarelli (2009), we can decompose:
$$
B_i = \underbrace{\bar{B}}_{\text{common}} + \underbrace{\tilde{B}_i}_{\text{country-specific}} + \underbrace{Z_i \gamma}_{\text{observed heterogeneity}}
$$
where:
- $\bar{B}$: Common component (global dynamics)
- $\tilde{B}_i$: Idiosyncratic deviation
- $Z_i$: Country characteristics (GDP per capita, openness, institutions)
```{r}
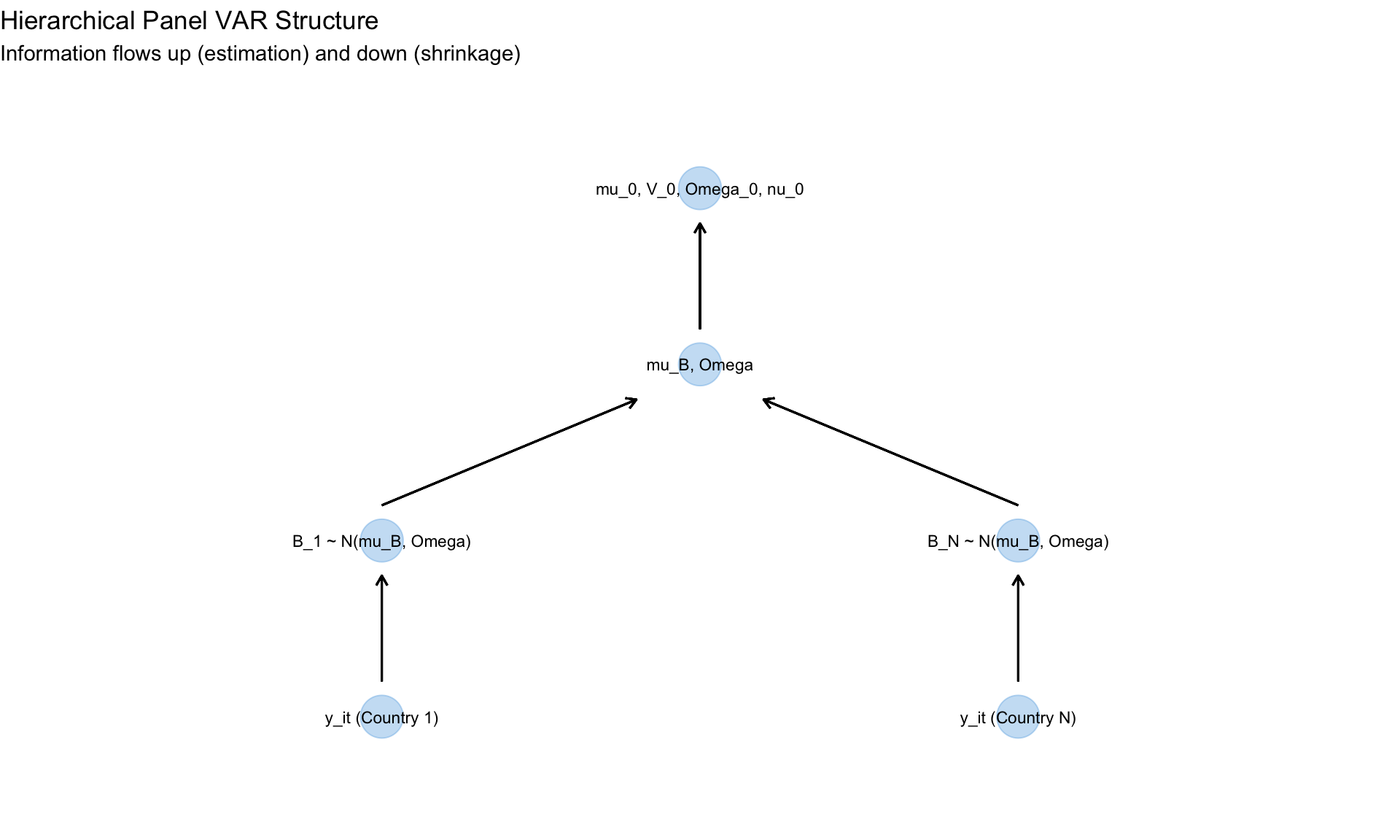
#| label: panel-var-setup
#| fig-cap: "Panel VAR structure for cross-country macro analysis"
# Illustrate panel VAR structure
panel_var_diagram <- tibble(
level = c("Data", "Data", "Country VAR", "Country VAR", "Hierarchical", "Hyperprior"),
component = c("y_it (Country 1)", "y_it (Country N)",
"B_1 ~ N(mu_B, Omega)", "B_N ~ N(mu_B, Omega)",
"mu_B, Omega", "mu_0, V_0, Omega_0, nu_0"),
x = c(1, 3, 1, 3, 2, 2),
y = c(1, 1, 2, 2, 3, 4)
)
ggplot(panel_var_diagram, aes(x = x, y = y, label = component)) +
geom_point(size = 10, color = "#3498db", alpha = 0.3) +
geom_text(size = 3) +
geom_segment(aes(x = 1, xend = 1, y = 1.2, yend = 1.8),
arrow = arrow(length = unit(0.2, "cm"))) +
geom_segment(aes(x = 3, xend = 3, y = 1.2, yend = 1.8),
arrow = arrow(length = unit(0.2, "cm"))) +
geom_segment(aes(x = 1, xend = 1.8, y = 2.2, yend = 2.8),
arrow = arrow(length = unit(0.2, "cm"))) +
geom_segment(aes(x = 3, xend = 2.2, y = 2.2, yend = 2.8),
arrow = arrow(length = unit(0.2, "cm"))) +
geom_segment(aes(x = 2, xend = 2, y = 3.2, yend = 3.8),
arrow = arrow(length = unit(0.2, "cm"))) +
xlim(0, 4) + ylim(0.5, 4.5) +
labs(title = "Hierarchical Panel VAR Structure",
subtitle = "Information flows up (estimation) and down (shrinkage)") +
theme_void()
```
## Gibbs Sampler for Panel VAR
```{r}
#| label: panel-var-gibbs
#| cache: true
# Simplified Panel VAR Gibbs sampler
# For K=2 variables, p=1 lag, demonstration purposes
panel_var_gibbs <- function(Y_list, p = 1, n_draw = 3000, n_burn = 1000,
lambda1 = 0.2) {
N <- length(Y_list) # number of countries
K <- ncol(Y_list[[1]]) # number of variables
M <- K * p + 1 # regressors per equation
# Build data matrices for each country
data_list <- map(Y_list, function(Y) {
T_full <- nrow(Y)
y <- Y[(p+1):T_full, , drop = FALSE]
X <- matrix(1, nrow = T_full - p, ncol = 1)
for (j in 1:p) {
X <- cbind(X, Y[(p+1-j):(T_full-j), , drop = FALSE])
}
list(y = y, X = X, T_eff = nrow(y))
})
# Hierarchical priors
# Level 2: mu_B ~ N(0, 10*I), Omega ~ IW(I, K+2)
mu_B_prior <- rep(0, M * K)
V_mu_prior <- diag(10, M * K)
Omega_prior <- diag(1, M * K)
nu_Omega_prior <- M * K + 2
# Storage
B_draws <- array(0, dim = c(M, K, N, n_draw))
mu_B_draws <- matrix(0, n_draw, M * K)
# Initialize
B_curr <- array(0, dim = c(M, K, N))
for (i in 1:N) {
B_curr[,,i] <- solve(crossprod(data_list[[i]]$X)) %*%
crossprod(data_list[[i]]$X, data_list[[i]]$y)
}
Sigma_curr <- lapply(1:N, function(i) {
U <- data_list[[i]]$y - data_list[[i]]$X %*% B_curr[,,i]
crossprod(U) / data_list[[i]]$T_eff
})
mu_B_curr <- apply(B_curr, c(1,2), mean) %>% as.vector()
Omega_curr <- diag(lambda1^2, M * K)
for (g in 1:(n_draw + n_burn)) {
# Block 1: Draw B_i | Sigma_i, mu_B, Omega for each country
for (i in 1:N) {
X_i <- data_list[[i]]$X
y_i <- data_list[[i]]$y
Sigma_inv <- solve(Sigma_curr[[i]])
Omega_inv <- solve(Omega_curr)
V_post_inv <- Omega_inv + kronecker(Sigma_inv, crossprod(X_i))
V_post <- solve(V_post_inv)
b_post <- V_post %*% (Omega_inv %*% mu_B_curr +
kronecker(Sigma_inv, t(X_i)) %*% as.vector(y_i))
B_curr[,,i] <- matrix(rmvnorm(1, b_post, V_post), M, K)
}
# Block 2: Draw Sigma_i | B_i for each country
for (i in 1:N) {
U <- data_list[[i]]$y - data_list[[i]]$X %*% B_curr[,,i]
S_post <- diag(K) + crossprod(U)
Sigma_curr[[i]] <- solve(rWishart(1, K + 2 + data_list[[i]]$T_eff,
solve(S_post))[,,1])
}
# Block 3: Draw mu_B | B_1,...,B_N, Omega
B_vec <- sapply(1:N, function(i) as.vector(B_curr[,,i]))
B_mean <- rowMeans(B_vec)
Omega_inv <- solve(Omega_curr)
V_mu_inv <- solve(V_mu_prior)
V_mu_post <- solve(N * Omega_inv + V_mu_inv)
m_mu_post <- V_mu_post %*% (N * Omega_inv %*% B_mean + V_mu_inv %*% mu_B_prior)
mu_B_curr <- as.vector(rmvnorm(1, m_mu_post, V_mu_post))
# Block 4: Draw Omega | B_1,...,B_N, mu_B
S_Omega <- Omega_prior
for (i in 1:N) {
b_i <- as.vector(B_curr[,,i])
S_Omega <- S_Omega + outer(b_i - mu_B_curr, b_i - mu_B_curr)
}
Omega_curr <- solve(rWishart(1, nu_Omega_prior + N, solve(S_Omega))[,,1])
# Store
if (g > n_burn) {
idx <- g - n_burn
B_draws[,,,idx] <- B_curr
mu_B_draws[idx,] <- mu_B_curr
}
if (g %% 500 == 0) cat("Draw", g, "of", n_draw + n_burn, "\n")
}
list(B = B_draws, mu_B = mu_B_draws, K = K, p = p, M = M, N = N)
}
# Generate synthetic panel VAR data
set.seed(123)
N_countries <- 10
T_obs <- 60
K <- 2
# Common VAR structure with heterogeneity
A_common <- matrix(c(0.7, 0.1, 0.05, 0.6), K, K)
Sigma_common <- matrix(c(1, 0.3, 0.3, 1), K, K)
Y_list <- map(1:N_countries, function(i) {
# Country-specific deviation
A_i <- A_common + matrix(rnorm(K^2, 0, 0.1), K, K)
Y <- matrix(0, T_obs, K)
Y[1, ] <- rnorm(K)
for (t in 2:T_obs) {
Y[t, ] <- Y[t-1, ] %*% t(A_i) + rmvnorm(1, sigma = Sigma_common)
}
Y
})
# Estimate panel VAR
cat("Estimating Panel VAR...\n")
pvar_result <- panel_var_gibbs(Y_list, p = 1, n_draw = 2000, n_burn = 500)
```
```{r}
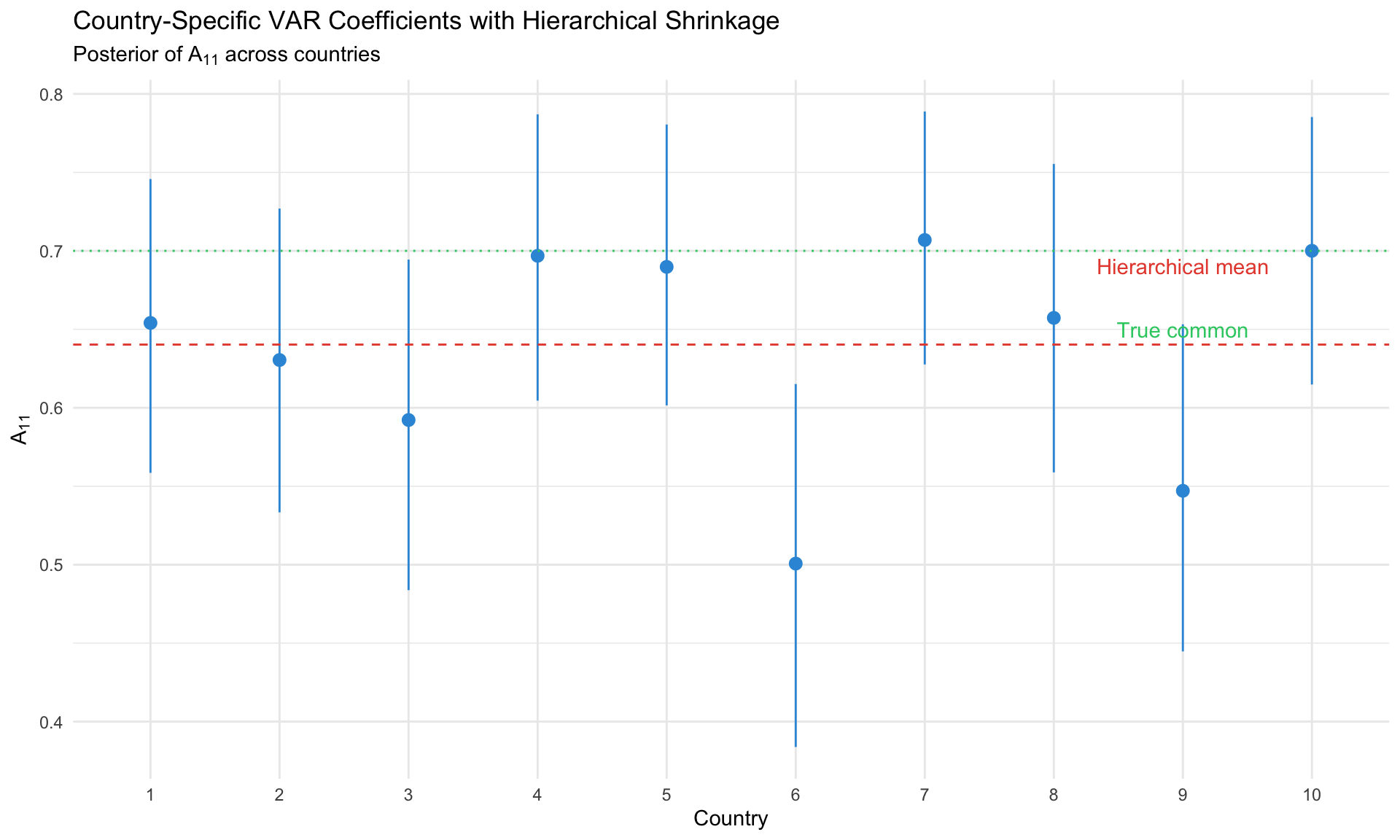
#| label: panel-var-results
#| fig-cap: "Panel VAR coefficient posteriors across countries"
# Extract A[1,1] coefficient for each country
A11_draws <- pvar_result$B[2, 1, , ] # position [2,1] is A[1,1] after constant
# Posterior summaries
A11_summary <- tibble(
country = 1:N_countries,
mean = apply(A11_draws, 1, mean),
lower = apply(A11_draws, 1, quantile, 0.16),
upper = apply(A11_draws, 1, quantile, 0.84)
)
# Add global mean
mu_A11 <- mean(pvar_result$mu_B[, 2]) # position 2 in vec(B) is A[1,1]
ggplot(A11_summary, aes(x = factor(country), y = mean)) +
geom_pointrange(aes(ymin = lower, ymax = upper), color = "#3498db") +
geom_hline(yintercept = mu_A11, linetype = "dashed", color = "#e74c3c") +
geom_hline(yintercept = A_common[1,1], linetype = "dotted", color = "#2ecc71") +
annotate("text", x = 9, y = mu_A11 + 0.05, label = "Hierarchical mean", color = "#e74c3c") +
annotate("text", x = 9, y = A_common[1,1] - 0.05, label = "True common", color = "#2ecc71") +
labs(title = "Country-Specific VAR Coefficients with Hierarchical Shrinkage",
subtitle = expression(paste("Posterior of ", A[11], " across countries")),
x = "Country", y = expression(A[11])) +
theme_minimal()
```
# Random Coefficients Models {.unnumbered}
## Swamy (1970) Random Coefficients
The classic frequentist approach:
$$
y_{it} = x_{it}'\beta_i + \varepsilon_{it}
$$
$$
\beta_i = \bar{\beta} + v_i, \quad v_i \sim N(0, \Omega)
$$
**Bayesian version** adds priors on $(\bar{\beta}, \Omega)$ and uses Gibbs sampling.
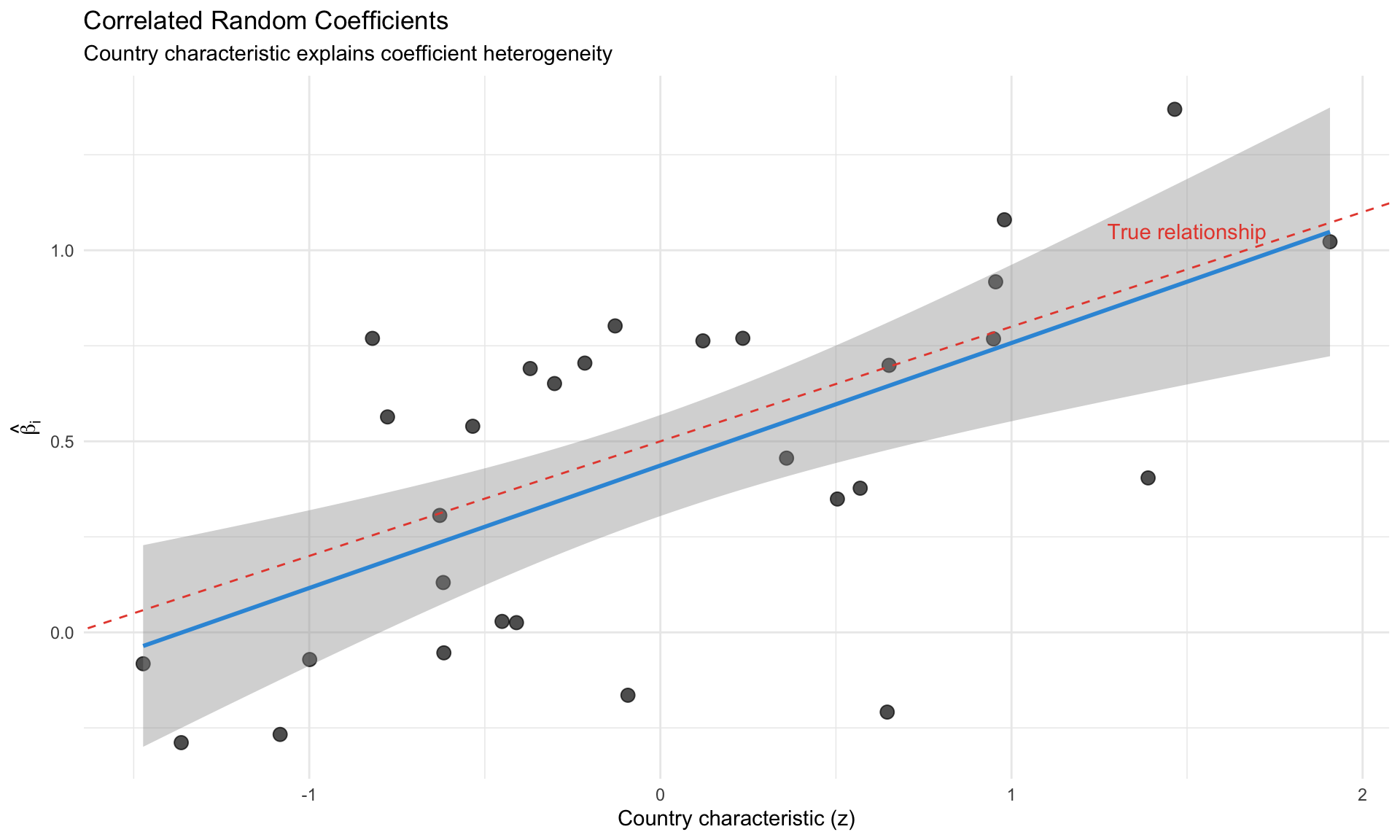
## Correlated Random Coefficients
Allow $\beta_i$ to correlate with observables:
$$
\beta_i = \bar{\beta} + Z_i'\gamma + v_i
$$
where $Z_i$ are country characteristics (GDP per capita, openness, institutions).
**Application**: Countries with higher institutional quality might have different Taylor rule coefficients.
```{r}
#| label: random-coef
#| fig-cap: "Random coefficients with observable heterogeneity"
# Simulate correlated random coefficients
N <- 30
T_obs <- 40
# Country characteristic
z_i <- rnorm(N) # e.g., log GDP per capita
# Coefficient depends on characteristic
gamma <- 0.3
beta_i <- 0.5 + gamma * z_i + rnorm(N, 0, 0.2)
# Generate panel
rc_data <- map_dfr(1:N, function(i) {
x <- rnorm(T_obs)
y <- beta_i[i] * x + rnorm(T_obs, 0, 1)
tibble(country = i, x = x, y = y, z = z_i[i])
})
# OLS by country
ols_rc <- rc_data %>%
group_by(country) %>%
summarise(beta_ols = coef(lm(y ~ x - 1))[1], z = first(z))
# Plot relationship
ggplot(ols_rc, aes(x = z, y = beta_ols)) +
geom_point(size = 3, alpha = 0.7) +
geom_smooth(method = "lm", se = TRUE, color = "#3498db") +
geom_abline(intercept = 0.5, slope = gamma, linetype = "dashed", color = "#e74c3c") +
annotate("text", x = 1.5, y = 0.5 + gamma * 1.5 + 0.1,
label = "True relationship", color = "#e74c3c") +
labs(title = "Correlated Random Coefficients",
subtitle = "Country characteristic explains coefficient heterogeneity",
x = "Country characteristic (z)", y = expression(hat(beta)[i])) +
theme_minimal()
```
# Cross-Sectional Dependence {.unnumbered}
## The Problem
Standard panel methods assume:
$$
E[\varepsilon_{it} \varepsilon_{js}] = 0 \quad \text{for } i \neq j
$$
But in macro panels, **global shocks** (oil prices, US monetary policy, COVID) create cross-sectional dependence.
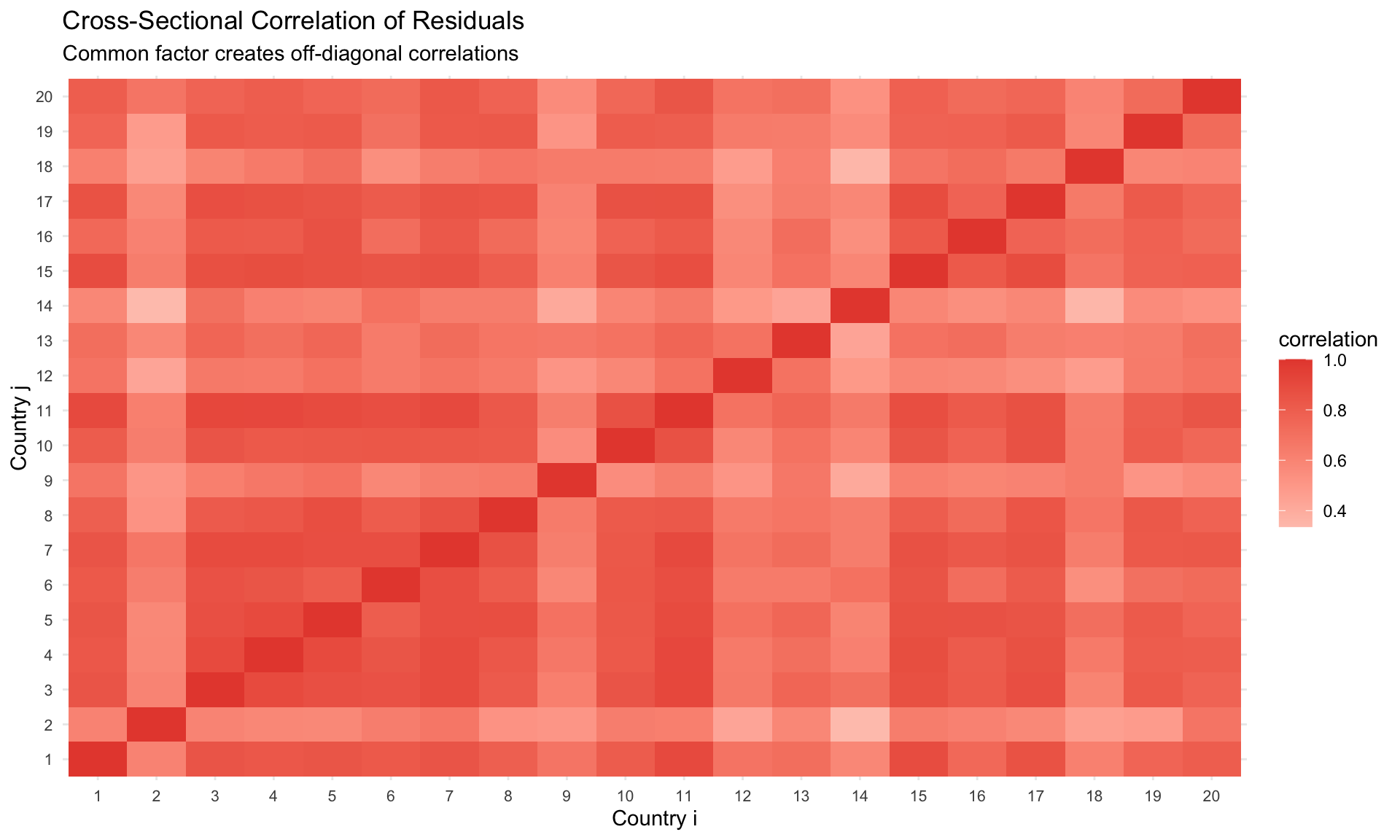
## Factor Structure Approach
Model the error as:
$$
\varepsilon_{it} = \lambda_i' f_t + e_{it}
$$
where:
- $f_t$: Common factors (K_f × 1)
- $\lambda_i$: Country-specific factor loadings
- $e_{it}$: Idiosyncratic error
### Interactive Fixed Effects (Bai 2009)
$$
y_{it} = x_{it}'\beta + \lambda_i' f_t + e_{it}
$$
Jointly estimates $\beta$, $\Lambda = [\lambda_1, \ldots, \lambda_N]'$, and $F = [f_1, \ldots, f_T]'$.
### Bayesian Factor Model
**Prior structure**:
$$
f_t | \Sigma_f \sim N(0, \Sigma_f), \quad \lambda_i \sim N(0, \Omega_\lambda)
$$
**Identification**: Impose lower-triangular structure on loadings (first K_f countries).
```{r}
#| label: factor-model
#| fig-cap: "Cross-sectional dependence via common factor"
# Simulate panel with common factor
N <- 20
T_obs <- 50
K_f <- 1 # one common factor
# Common factor (e.g., global financial cycle)
f_t <- arima.sim(model = list(ar = 0.8), n = T_obs) %>% as.vector()
# Country loadings
lambda_i <- rnorm(N, mean = 0.5, sd = 0.3)
# Generate correlated errors
errors <- outer(f_t, lambda_i) + matrix(rnorm(T_obs * N, 0, 0.5), T_obs, N)
# True coefficient
beta_true <- 0.8
# Generate data
factor_data <- map_dfr(1:N, function(i) {
x <- rnorm(T_obs)
y <- beta_true * x + errors[, i]
tibble(country = i, t = 1:T_obs, x = x, y = y, factor = f_t)
})
# Show cross-correlation of residuals
resids <- factor_data %>%
group_by(country) %>%
mutate(resid = residuals(lm(y ~ x))) %>%
select(country, t, resid) %>%
pivot_wider(names_from = country, values_from = resid, names_prefix = "c")
# Correlation matrix of residuals
resid_mat <- as.matrix(resids[, -1])
cor_mat <- cor(resid_mat)
# Visualize
cor_df <- expand_grid(i = 1:N, j = 1:N) %>%
mutate(correlation = as.vector(cor_mat))
ggplot(cor_df, aes(x = factor(i), y = factor(j), fill = correlation)) +
geom_tile() +
scale_fill_gradient2(low = "#3498db", mid = "white", high = "#e74c3c", midpoint = 0) +
labs(title = "Cross-Sectional Correlation of Residuals",
subtitle = "Common factor creates off-diagonal correlations",
x = "Country i", y = "Country j") +
theme_minimal() +
theme(axis.text = element_text(size = 8))
```
## Bayesian Factor-Augmented Panel VAR
Combine hierarchical panel VAR with factor structure:
$$
y_{it} = c_i + A_i y_{i,t-1} + \Gamma_i f_t + u_{it}
$$
where $f_t$ is a vector of global factors (extracted or observed).
**Application in macro**: Include Global Financial Cycle (GFC) factor from Miranda-Agrippino & Rey.
# Python Implementation {.unnumbered}
For larger panels or more complex hierarchical structures, Python with NumPy/SciPy provides flexibility.
```python
# Hierarchical Panel Model in Python
import numpy as np
from scipy import stats
from scipy.linalg import solve, cholesky
def hierarchical_panel_gibbs(y_list, X_list, n_draw=5000, n_burn=1000):
"""
Gibbs sampler for hierarchical panel regression.
y_list: list of (T_i,) arrays - outcomes by country
X_list: list of (T_i, K) arrays - regressors by country
"""
N = len(y_list)
K = X_list[0].shape[1]
# Priors
mu0 = np.zeros(K)
V0 = np.eye(K) * 10
V0_inv = np.linalg.inv(V0)
Omega0 = np.eye(K)
nu0 = K + 2
a0, b0 = 0.01, 0.01 # for sigma^2
# Storage
beta_draws = np.zeros((n_draw, N, K))
mu_draws = np.zeros((n_draw, K))
Omega_draws = np.zeros((n_draw, K, K))
# Initialize
beta_curr = np.zeros((N, K))
for i in range(N):
beta_curr[i] = np.linalg.lstsq(X_list[i], y_list[i], rcond=None)[0]
sigma2_curr = np.ones(N)
mu_curr = beta_curr.mean(axis=0)
Omega_curr = np.cov(beta_curr.T) + 0.01 * np.eye(K)
for g in range(n_draw + n_burn):
Omega_inv = np.linalg.inv(Omega_curr)
# Block 1: Draw beta_i | rest
for i in range(N):
X_i, y_i = X_list[i], y_list[i]
XtX = X_i.T @ X_i
Xty = X_i.T @ y_i
V_post_inv = XtX / sigma2_curr[i] + Omega_inv
V_post = np.linalg.inv(V_post_inv)
m_post = V_post @ (Xty / sigma2_curr[i] + Omega_inv @ mu_curr)
beta_curr[i] = np.random.multivariate_normal(m_post, V_post)
# Block 2: Draw sigma2_i | rest
for i in range(N):
resid = y_list[i] - X_list[i] @ beta_curr[i]
T_i = len(y_list[i])
a_post = a0 + T_i / 2
b_post = b0 + np.sum(resid**2) / 2
sigma2_curr[i] = 1 / np.random.gamma(a_post, 1/b_post)
# Block 3: Draw mu | rest
V_mu_post = np.linalg.inv(N * Omega_inv + V0_inv)
m_mu_post = V_mu_post @ (N * Omega_inv @ beta_curr.mean(axis=0) +
V0_inv @ mu0)
mu_curr = np.random.multivariate_normal(m_mu_post, V_mu_post)
# Block 4: Draw Omega | rest (Inverse Wishart)
S_post = Omega0.copy()
for i in range(N):
diff = beta_curr[i] - mu_curr
S_post += np.outer(diff, diff)
# Draw from Inverse Wishart
df_post = nu0 + N
Omega_curr = stats.invwishart.rvs(df=df_post, scale=S_post)
# Store
if g >= n_burn:
idx = g - n_burn
beta_draws[idx] = beta_curr
mu_draws[idx] = mu_curr
Omega_draws[idx] = Omega_curr
return {
'beta': beta_draws,
'mu': mu_draws,
'Omega': Omega_draws
}
# Example usage:
# result = hierarchical_panel_gibbs(y_list, X_list)
# beta_posterior_mean = result['beta'].mean(axis=0)
```
# Model Comparison for Panel Models {.unnumbered}
## Marginal Likelihood
For hierarchical models, the marginal likelihood integrates over all parameters:
$$
p(Y | M) = \int p(Y | \theta) p(\theta | M) d\theta
$$
**For linear hierarchical models**: Often tractable analytically.
**For complex models**: Use bridge sampling or the Chib (1995) method.
## DIC and WAIC
| Criterion | Formula | Use |
|-----------|---------|-----|
| DIC | $\bar{D} + p_D$ | Effective parameters penalty |
| WAIC | Pointwise predictive density | Better for hierarchical models |
| LOO-CV | Leave-one-out cross-validation | Gold standard but expensive |
```{r}
#| label: dic-calculation
# DIC calculation for hierarchical model
calculate_dic <- function(hier_result, data_hier) {
countries <- unique(data_hier$country)
N <- length(countries)
n_draw <- dim(hier_result$beta)[3]
# Compute log-likelihood at each draw
log_lik_draws <- numeric(n_draw)
for (g in 1:n_draw) {
ll <- 0
for (i in 1:N) {
d <- filter(data_hier, country == i)
beta_g <- hier_result$beta[i, 1, g]
sigma2_g <- hier_result$sigma2[g, i]
resid <- d$y - d$x * beta_g
ll <- ll + sum(dnorm(resid, 0, sqrt(sigma2_g), log = TRUE))
}
log_lik_draws[g] <- ll
}
# DIC components
D_bar <- -2 * mean(log_lik_draws) # posterior mean deviance
# Deviance at posterior mean
ll_at_mean <- 0
for (i in 1:N) {
d <- filter(data_hier, country == i)
beta_mean <- mean(hier_result$beta[i, 1, ])
sigma2_mean <- mean(hier_result$sigma2[, i])
resid <- d$y - d$x * beta_mean
ll_at_mean <- ll_at_mean + sum(dnorm(resid, 0, sqrt(sigma2_mean), log = TRUE))
}
D_theta_bar <- -2 * ll_at_mean
# Effective parameters
p_D <- D_bar - D_theta_bar
# DIC
DIC <- D_bar + p_D
list(DIC = DIC, D_bar = D_bar, p_D = p_D)
}
dic_result <- calculate_dic(hier_result, data_hier)
cat("DIC:", round(dic_result$DIC, 1), "\n")
cat("Effective parameters (p_D):", round(dic_result$p_D, 1), "\n")
```
# Practical Guidance {.unnumbered}
## When to Use Hierarchical vs. Standard Panel
| Situation | Recommendation |
|-----------|----------------|
| Small T per country (< 30) | Hierarchical (shrinkage helps) |
| Large N, moderate T | Hierarchical (efficient) |
| Focus on average effect | Either works |
| Focus on heterogeneity | Hierarchical (full posterior) |
| Strong prior beliefs | Bayesian hierarchical |
## Implementation Checklist
1. **Check MCMC convergence**: Trace plots, R-hat, effective sample size
2. **Assess shrinkage**: Compare hierarchical to OLS estimates
3. **Test homogeneity**: Is $\Omega$ small relative to coefficient magnitudes?
4. **Cross-sectional dependence**: Test residuals, consider factors
5. **Model comparison**: DIC/WAIC across specifications
## Common Pitfalls
::: {.callout-warning}
## Watch Out For
1. **Over-shrinkage**: Very tight hyperpriors force all countries to look identical
2. **Under-shrinkage**: Vague hyperpriors fail to borrow strength
3. **Ignoring cross-sectional dependence**: Biases standard errors downward
4. **Too many parameters**: K-variable panel VAR with p lags has $N \times K \times (Kp + 1)$ parameters
5. **Improper mixing**: Hierarchical models can have slow mixing — increase draws
:::
# Key References {.unnumbered}
## Hierarchical Panel Models
- **Gelman & Hill (2007)** *Data Analysis Using Regression and Multilevel/Hierarchical Models* — Accessible introduction
- **Swamy (1970)** "Efficient Inference in a Random Coefficient Regression Model" Econometrica — Classic random coefficients
- **Hsiao (2014)** *Analysis of Panel Data* — Comprehensive textbook
## Bayesian Panel VAR
- **Canova & Ciccarelli (2009)** "Estimating Multicountry VAR Models" IER — Foundational paper
- **Canova & Ciccarelli (2013)** "Panel Vector Autoregressive Models: A Survey" — Survey chapter
- **Jarocinski (2010)** "Responses to Monetary Policy Shocks in the East and the West of Europe" JAE — Application
## Cross-Sectional Dependence
- **Bai (2009)** "Panel Data Models with Interactive Fixed Effects" Econometrica — Factor structure
- **Pesaran (2006)** "Estimation and Inference in Large Heterogeneous Panels" Econometrica — CCE estimator
- **Chudik & Pesaran (2015)** "Common Correlated Effects Estimation of Heterogeneous Dynamic Panel Data Models" JoE
## Software
- **R**: `brms` (Bürkner), `lme4` (Bates et al.), custom Gibbs samplers
- **Python**: `PyMC`, NumPy/SciPy for manual MCMC
- **Stan**: `rstan`/`pystan` for complex hierarchical models
# Summary {.unnumbered}
| Concept | Key Insight |
|---------|-------------|
| **Hierarchical models** | Pool information while allowing heterogeneity |
| **Shrinkage** | Data-poor countries borrow from data-rich |
| **Panel VAR** | Country-specific dynamics with common structure |
| **Cross-sectional dependence** | Global factors create correlations |
| **Gibbs sampling** | Natural framework for hierarchical models |
::: {.callout-tip}
## For Your Research
With N = 46 countries and T = 84 quarters:
- Hierarchical shrinkage is valuable for EM countries with noisier data
- Panel VAR can reveal heterogeneity in policy transmission
- Factor structures capture global shocks (GFC, COVID)
- Always check cross-sectional correlation in residuals
:::