---
title: "Synthetic Control Methods"
subtitle: "Data-Driven Counterfactuals for Comparative Case Studies"
---
```{r}
#| label: setup
#| include: false
knitr::opts_chunk$set(
echo = TRUE, warning = FALSE, message = FALSE,
fig.width = 10, fig.height = 6, fig.align = "center"
)
library(dplyr)
library(tidyr)
library(ggplot2)
theme_set(theme_minimal(base_size = 13))
set.seed(42)
```
# When DiD Fails {.unnumbered}
Difference-in-differences requires **parallel trends**—the assumption that treated and control groups would have followed similar paths absent treatment. But what happens when:
- You have only **one treated unit**?
- No control unit has trends parallel to the treated unit?
- The treated unit is unique (e.g., West Germany, California, a specific country)?
**Synthetic Control Methods (SCM)** solve this by constructing a weighted combination of untreated units that together replicate the treated unit's pre-treatment trajectory. Instead of assuming parallel trends, SCM **matches** them.
## The Core Insight
| Method | Comparison Unit | Key Assumption |
|--------|----------------|----------------|
| DiD | Actual control group | Parallel trends |
| SCM | **Synthetic** control (weighted donors) | Weights match pre-treatment outcomes |
SCM is transparent: you see exactly which units contribute to the counterfactual and with what weights.
# The SCM Framework {.unnumbered}
## Abadie, Diamond & Hainmueller (2010, 2015)
### Setup and Notation
Following the foundational papers:
$$
\begin{aligned}
&\text{Treated unit: } i = 1 \\
&\text{Donor pool: } i = 2, \ldots, J+1 \\
&\text{Pre-treatment: } t = 1, \ldots, T_0 \\
&\text{Post-treatment: } t = T_0+1, \ldots, T
\end{aligned}
$$
We want to estimate the treatment effect:
$$
\tau_{1t} = Y_{1t}(1) - Y_{1t}(0) \quad \text{for } t > T_0
$$
We observe $Y_{1t}(1)$ (the actual outcome under treatment). The challenge is estimating $Y_{1t}(0)$—what would have happened without treatment.
### The Synthetic Control Estimator
Find weights $W^* = (w_2, \ldots, w_{J+1})'$ that solve:
$$
\min_W \|X_1 - X_0 W\|_V \quad \text{subject to } w_j \geq 0, \sum_{j=2}^{J+1} w_j = 1
$$
where:
- $X_1$ = vector of pre-treatment characteristics of treated unit
- $X_0$ = matrix of pre-treatment characteristics of donor units
- $V$ = diagonal weight matrix (predictor importance)
- Constraints ensure weights are non-negative and sum to one (convex combination)
**The synthetic control outcome**:
$$
\hat{Y}_{1t}(0) = \sum_{j=2}^{J+1} w_j^* Y_{jt}
$$
**The treatment effect estimate**:
$$
\hat{\tau}_{1t} = Y_{1t} - \sum_{j=2}^{J+1} w_j^* Y_{jt}
$$
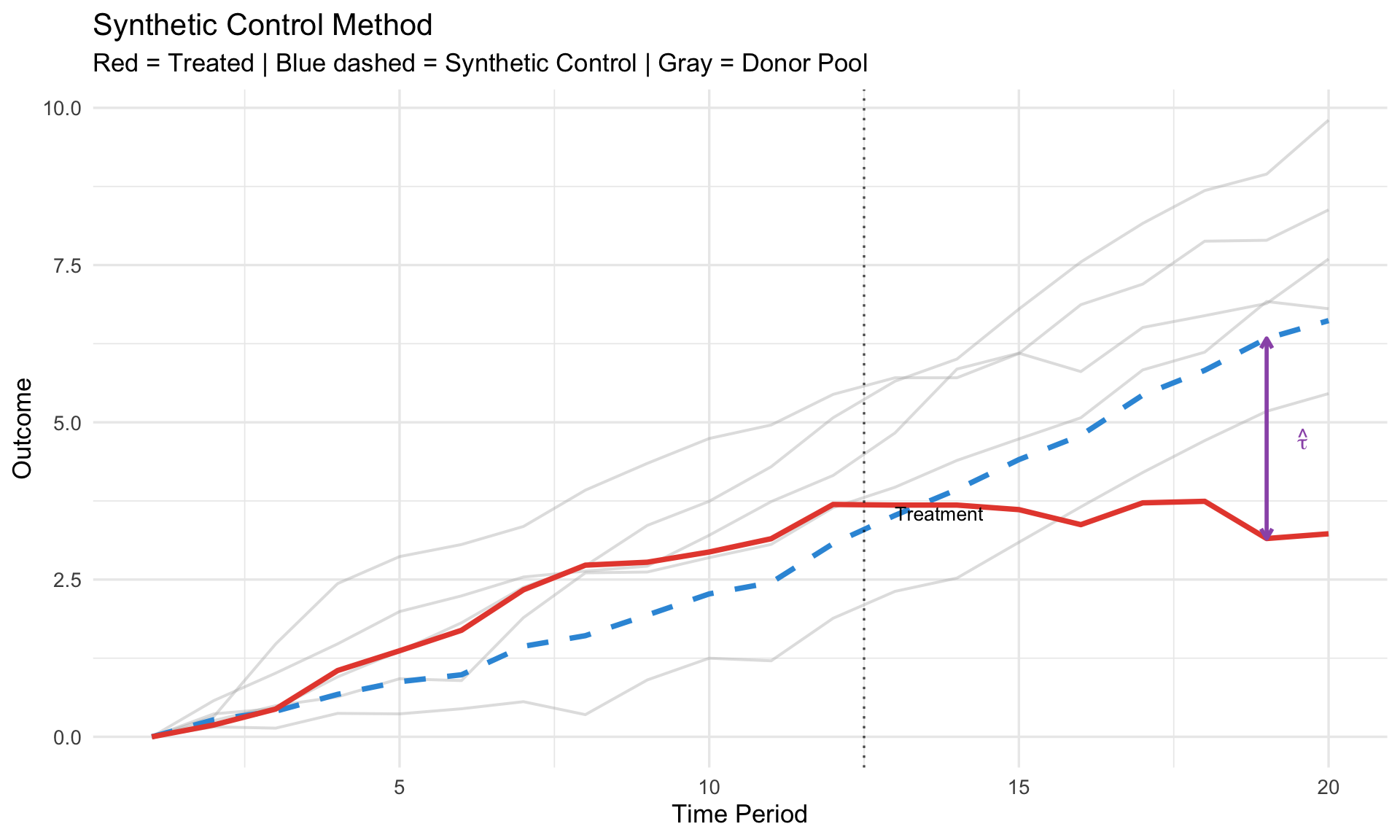
```{r}
#| label: scm-visual
#| fig-cap: "Synthetic control: construct counterfactual from weighted donors"
# Simulate SCM scenario
set.seed(123)
T_total <- 20
T0 <- 12 # Treatment after period 12
# Treated unit trajectory
treated_base <- cumsum(c(0, rnorm(T0-1, 0.3, 0.2)))
treated_post <- treated_base[T0] + cumsum(rnorm(T_total - T0, -0.1, 0.25))
treated <- c(treated_base, treated_post)
# Donor units (5 potential controls)
donors <- lapply(1:5, function(i) {
base <- cumsum(c(0, rnorm(T_total-1, 0.25 + 0.05*i, 0.3)))
data.frame(time = 1:T_total, unit = paste("Donor", i), y = base)
}) %>% bind_rows()
# Optimal weights (illustrative - found to match pre-treatment)
weights <- c(0.45, 0.30, 0.15, 0.10, 0.00)
# Construct synthetic control
synthetic <- donors %>%
mutate(weight = case_when(
unit == "Donor 1" ~ weights[1],
unit == "Donor 2" ~ weights[2],
unit == "Donor 3" ~ weights[3],
unit == "Donor 4" ~ weights[4],
TRUE ~ weights[5]
)) %>%
group_by(time) %>%
summarize(y = sum(y * weight), .groups = "drop")
# Combine for plotting
treated_df <- data.frame(time = 1:T_total, y = treated, unit = "Treated")
ggplot() +
# Donor units (faded)
geom_line(data = donors, aes(x = time, y = y, group = unit),
color = "gray70", alpha = 0.4, linewidth = 0.7) +
# Synthetic control
geom_line(data = synthetic, aes(x = time, y = y),
color = "#3498db", linewidth = 1.3, linetype = "dashed") +
# Treated unit
geom_line(data = treated_df, aes(x = time, y = y),
color = "#e74c3c", linewidth = 1.3) +
# Treatment line
geom_vline(xintercept = T0 + 0.5, linetype = "dotted", alpha = 0.7) +
annotate("text", x = T0 + 1, y = max(treated) * 0.95,
label = "Treatment", hjust = 0, size = 3.5) +
# Gap annotation
annotate("segment", x = T_total - 1, xend = T_total - 1,
y = treated[T_total-1], yend = synthetic$y[T_total-1],
arrow = arrow(ends = "both", length = unit(0.08, "inches")),
color = "#9b59b6", linewidth = 1) +
annotate("text", x = T_total - 0.5,
y = mean(c(treated[T_total-1], synthetic$y[T_total-1])),
label = expression(hat(tau)), hjust = 0, size = 4, color = "#9b59b6") +
labs(title = "Synthetic Control Method",
subtitle = "Red = Treated | Blue dashed = Synthetic Control | Gray = Donor Pool",
x = "Time Period", y = "Outcome") +
theme(legend.position = "none")
```
## Choosing Predictors ($X$)
The predictor matrix $X$ typically includes:
| Predictor Type | Examples | Rationale |
|----------------|----------|-----------|
| **Pre-treatment outcomes** | $Y_{1,T_0}, Y_{1,T_0-1}, \ldots$ | Forces trajectory matching |
| **Economic fundamentals** | GDP growth, inflation, trade openness | Structural similarity |
| **Demographics** | Population, urbanization | Background comparability |
| **Policy variables** | Tax rates, regulations | Institutional similarity |
::: {.callout-important}
## Include Pre-Treatment Outcomes
Abadie (2021) emphasizes: always include **multiple pre-treatment outcomes** as predictors. This forces the synthetic control to track the actual trajectory, not just match averages.
:::
## The Nested Optimization
In practice, SCM solves two nested problems:
**Inner problem** (given predictor weights $V$):
$$
W^*(V) = \arg\min_W (X_1 - X_0 W)' V (X_1 - X_0 W)
$$
**Outer problem** (choose $V$ to minimize pre-treatment fit):
$$
V^* = \arg\min_V \sum_{t=1}^{T_0} \left(Y_{1t} - \sum_{j} w_j^*(V) Y_{jt}\right)^2
$$
This ensures the weights are chosen to match the **outcome trajectory**, not just predictor averages.
# Inference: Permutation Tests {.unnumbered}
With one treated unit, standard inference doesn't apply. SCM uses **permutation-based inference**.
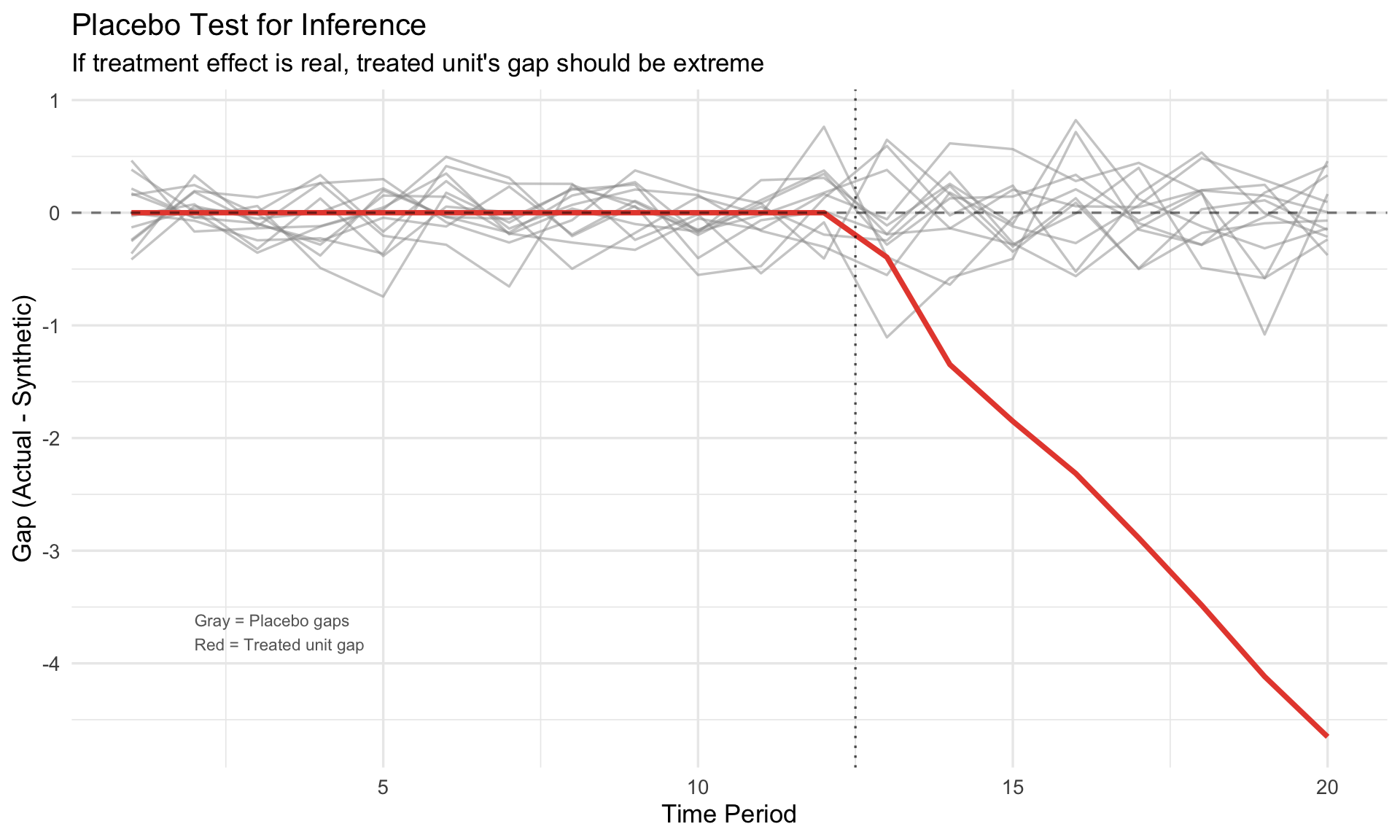
## The Placebo Test
**Idea**: Apply SCM to each donor unit as if it were treated. If the treated unit's effect is real, its gap should be unusually large compared to placebo gaps.
**Procedure**:
1. Estimate SCM for treated unit → get gap $\hat{\tau}_{1t}$
2. For each donor $j = 2, \ldots, J+1$:
- Pretend unit $j$ is treated (remove from donor pool)
- Construct synthetic control from remaining units
- Calculate placebo gap $\hat{\tau}_{jt}$
3. Compare: Is $|\hat{\tau}_{1t}|$ extreme relative to $\{|\hat{\tau}_{jt}|\}$?
**P-value**:
$$
p = \frac{\#\{j : |\hat{\tau}_j| \geq |\hat{\tau}_1|\}}{J + 1}
$$
```{r}
#| label: placebo-test
#| fig-cap: "Placebo test: treated unit gap vs. placebo distribution"
# Simulate placebo test
set.seed(789)
n_placebo <- 12
# Treated unit has real effect
treated_gap <- c(rep(0, T0), cumsum(rnorm(T_total - T0, -0.5, 0.2)))
# Placebo gaps (no real effect)
placebo_gaps <- lapply(1:n_placebo, function(i) {
gap <- c(rnorm(T0, 0, 0.25), rnorm(T_total - T0, 0, 0.35))
data.frame(time = 1:T_total, gap = gap, unit = paste("Placebo", i))
}) %>% bind_rows()
# Combine
all_gaps <- bind_rows(
data.frame(time = 1:T_total, gap = treated_gap, unit = "Treated"),
placebo_gaps
)
ggplot(all_gaps, aes(x = time, y = gap, group = unit)) +
geom_line(data = filter(all_gaps, unit != "Treated"),
color = "gray60", alpha = 0.5, linewidth = 0.6) +
geom_line(data = filter(all_gaps, unit == "Treated"),
color = "#e74c3c", linewidth = 1.3) +
geom_hline(yintercept = 0, linetype = "dashed", alpha = 0.5) +
geom_vline(xintercept = T0 + 0.5, linetype = "dotted", alpha = 0.7) +
annotate("text", x = 2, y = min(all_gaps$gap) * 0.8,
label = "Gray = Placebo gaps\nRed = Treated unit gap",
hjust = 0, size = 3, color = "gray40") +
labs(title = "Placebo Test for Inference",
subtitle = "If treatment effect is real, treated unit's gap should be extreme",
x = "Time Period", y = "Gap (Actual - Synthetic)")
```
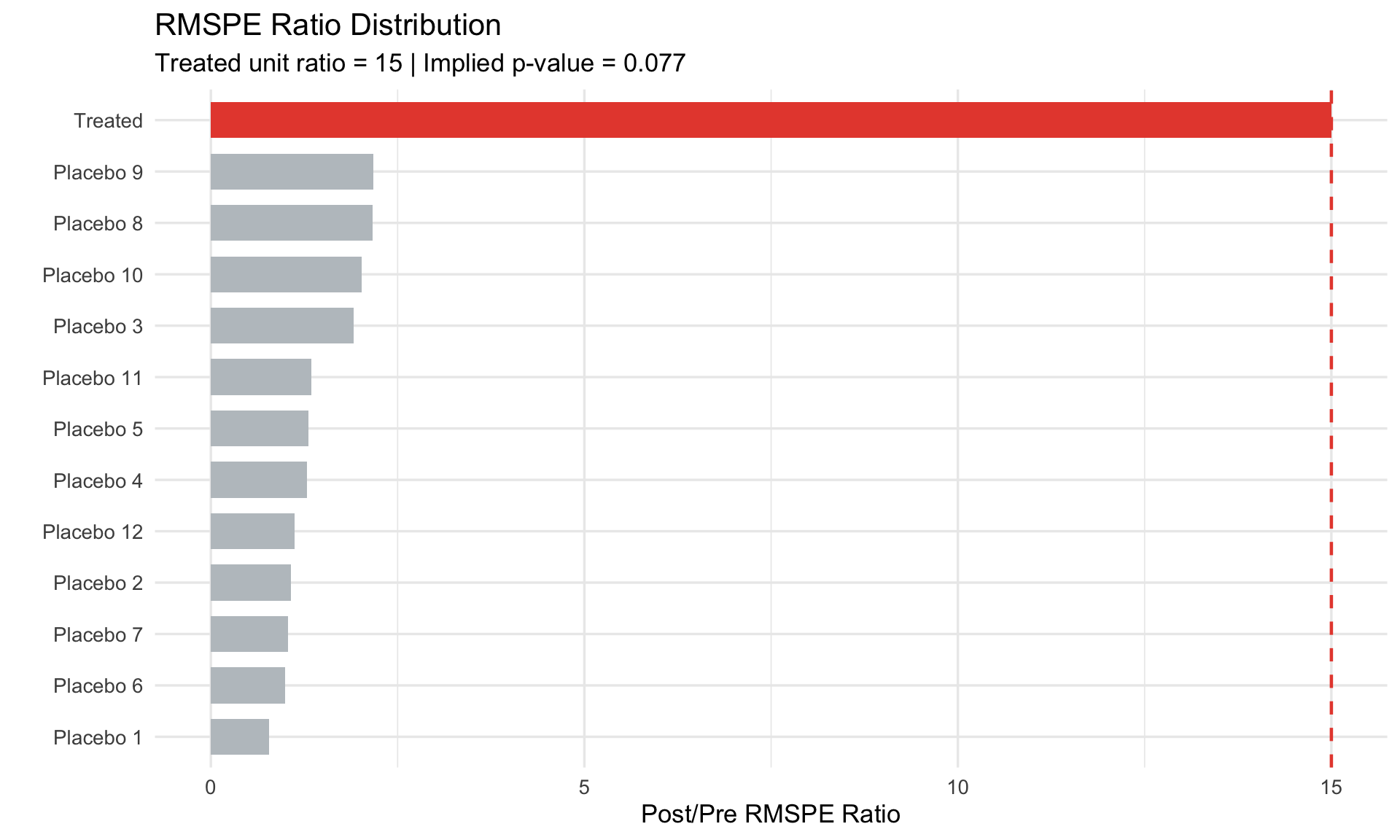
## RMSPE Ratio
To account for **pre-treatment fit quality**, use the RMSPE ratio:
$$
\text{Ratio}_j = \frac{\text{RMSPE}_{j,\text{post}}}{\text{RMSPE}_{j,\text{pre}}}
$$
where:
$$
\text{RMSPE}_{j,\text{pre}} = \sqrt{\frac{1}{T_0} \sum_{t=1}^{T_0} (Y_{jt} - \hat{Y}_{jt}^{\text{synth}})^2}
$$
**Rationale**: A unit with poor pre-treatment fit may have large post-treatment gaps just from noise. The ratio normalizes by pre-treatment fit quality.
```{r}
#| label: rmspe-ratio
#| fig-cap: "RMSPE ratio: normalizing by pre-treatment fit"
# Calculate RMSPE ratios
rmspe_pre <- c(0.12, runif(n_placebo, 0.15, 0.6)) # Treated has good fit
rmspe_post <- c(1.8, runif(n_placebo, 0.2, 1.2)) # Treated has large post gap
rmspe_df <- data.frame(
unit = c("Treated", paste("Placebo", 1:n_placebo)),
rmspe_pre = rmspe_pre,
rmspe_post = rmspe_post
) %>%
mutate(
ratio = rmspe_post / rmspe_pre,
is_treated = unit == "Treated"
) %>%
arrange(desc(ratio))
# P-value
p_val <- mean(rmspe_df$ratio >= rmspe_df$ratio[rmspe_df$unit == "Treated"])
ggplot(rmspe_df, aes(x = reorder(unit, ratio), y = ratio, fill = is_treated)) +
geom_col(width = 0.7) +
geom_hline(yintercept = rmspe_df$ratio[rmspe_df$unit == "Treated"],
linetype = "dashed", color = "#e74c3c", linewidth = 0.8) +
coord_flip() +
scale_fill_manual(values = c("FALSE" = "#bdc3c7", "TRUE" = "#e74c3c")) +
labs(title = "RMSPE Ratio Distribution",
subtitle = paste0("Treated unit ratio = ",
round(rmspe_df$ratio[rmspe_df$unit == "Treated"], 1),
" | Implied p-value = ", round(p_val, 3)),
x = "", y = "Post/Pre RMSPE Ratio") +
theme(legend.position = "none")
```
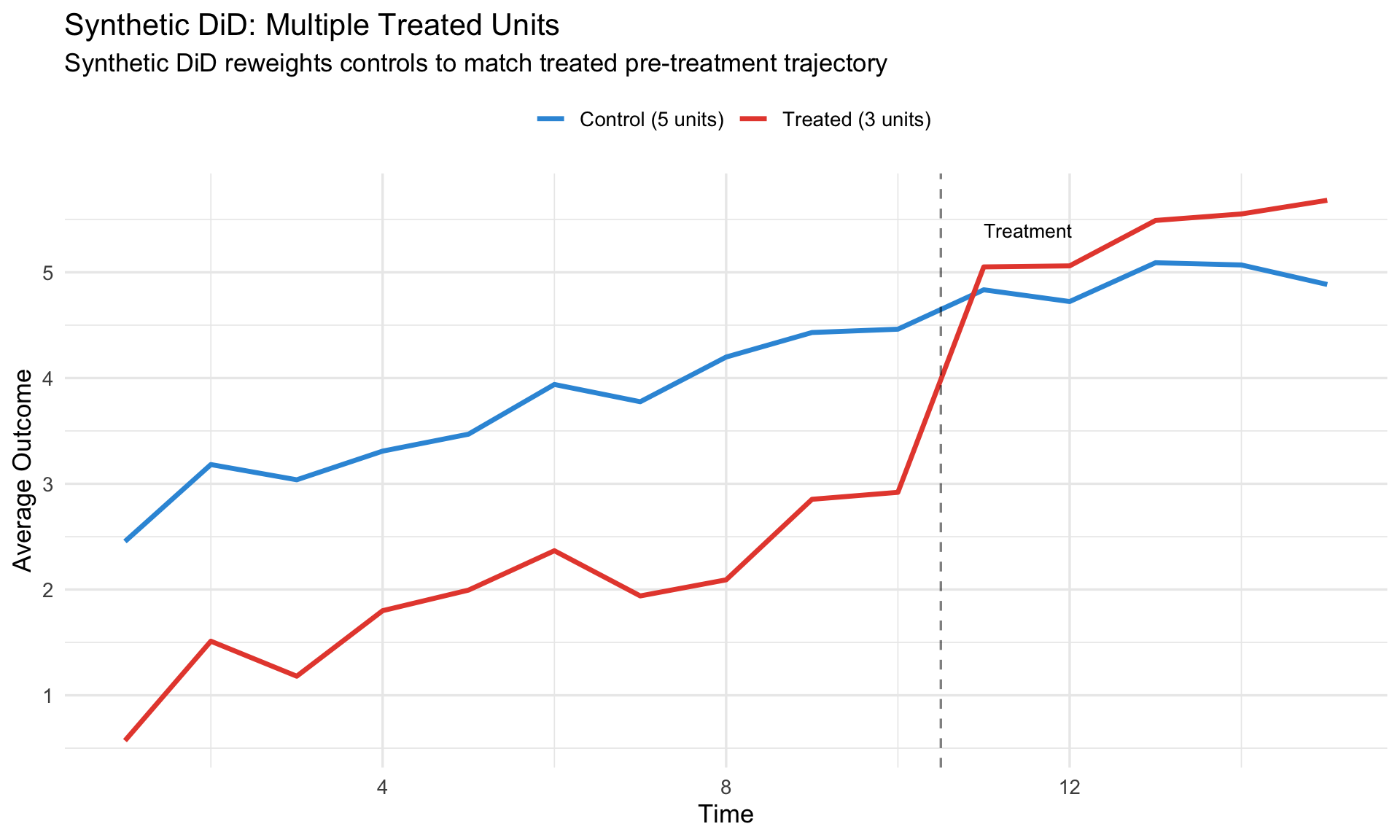
# Synthetic Difference-in-Differences {.unnumbered}
## Arkhangelsky et al. (2021)
**Synthetic DiD** combines the strengths of SCM and DiD:
| Method | Handles Multiple Treated? | Requires Parallel Trends? |
|--------|---------------------------|---------------------------|
| DiD | Yes | Yes |
| SCM | No (designed for one) | No (matches trajectories) |
| **Synthetic DiD** | **Yes** | **No** |
### The Estimator
Synthetic DiD finds:
1. **Unit weights** $\omega_i$ (like SCM) — make controls look like treated pre-treatment
2. **Time weights** $\lambda_t$ (novel) — emphasize pre-treatment periods that predict post-treatment
$$
\hat{\tau}^{sdid} = \left(\bar{Y}_{tr,post}^{\omega} - \bar{Y}_{co,post}^{\omega}\right) - \left(\bar{Y}_{tr,pre}^{\lambda} - \bar{Y}_{co,pre}^{\omega,\lambda}\right)
$$
The optimization:
$$
\min_{\omega, \lambda} \sum_{i: \text{control}} \sum_{t \leq T_0} \left(Y_{it} - \lambda_t - \omega_i - \mu\right)^2 + \text{regularization}
$$
### Advantages
1. **Works with multiple treated units** (unlike classic SCM)
2. **Doesn't require exact pre-treatment match** (unlike SCM)
3. **Doesn't require parallel trends** (unlike DiD)
4. **Doubly robust**: consistent if either unit OR time weights are correct
```{r}
#| label: sdid-comparison
#| fig-cap: "Synthetic DiD handles multiple treated units"
# Illustrate multiple treated units scenario
sdid_data <- expand.grid(
time = 1:15,
unit = 1:8
) %>%
mutate(
treated = unit <= 3,
post = time > 10,
treatment = treated & post,
# Generate outcomes
unit_fe = unit * 0.4,
time_trend = 0.2 * time,
tau = ifelse(treatment, 2, 0),
y = unit_fe + time_trend + tau + rnorm(n(), 0, 0.4),
group = ifelse(treated, "Treated (3 units)", "Control (5 units)")
)
# Aggregate by group
agg_data <- sdid_data %>%
group_by(time, group) %>%
summarize(y = mean(y), .groups = "drop")
ggplot(agg_data, aes(x = time, y = y, color = group)) +
geom_line(linewidth = 1.2) +
geom_vline(xintercept = 10.5, linetype = "dashed", alpha = 0.5) +
scale_color_manual(values = c("Control (5 units)" = "#3498db",
"Treated (3 units)" = "#e74c3c")) +
annotate("text", x = 11, y = max(agg_data$y) * 0.95,
label = "Treatment", hjust = 0, size = 3.5) +
labs(title = "Synthetic DiD: Multiple Treated Units",
subtitle = "Synthetic DiD reweights controls to match treated pre-treatment trajectory",
x = "Time", y = "Average Outcome",
color = "") +
theme(legend.position = "top")
```
# Implementation in R {.unnumbered}
## Using `augsynth` Package
The `augsynth` package by Ben-Michael et al. implements **augmented synthetic control**, which combines SCM with outcome modeling (Ridge regression) for improved finite-sample performance.
```r
library(augsynth)
# Basic synthetic control
syn <- augsynth(
outcome ~ treatment | covariate1 + covariate2, # outcome ~ treatment | predictors
unit = unit_id,
time = time_id,
data = panel_data,
t_int = treatment_time, # when treatment begins
progfunc = "Ridge" # augmentation: "None", "Ridge", "EN"
)
# Results
summary(syn)
plot(syn)
# Extract weights
syn$weights
```
### Key Arguments
| Argument | Description |
|----------|-------------|
| `progfunc` | Augmentation method: `"None"` (pure SCM), `"Ridge"`, `"EN"` (elastic net) |
| `t_int` | Treatment time (integer or date) |
| `fixedeff` | Include unit fixed effects in augmentation? |
### Multiple Treated Units with `multisynth`
```r
# When multiple units are treated (possibly at different times)
multi_syn <- multisynth(
outcome ~ treatment | covariates,
unit = unit_id,
time = time_id,
data = panel_data,
n_leads = 8, # post-treatment horizons to estimate
n_lags = 4 # pre-treatment periods to match
)
summary(multi_syn)
plot(multi_syn)
```
## Using `synthdid` Package
The `synthdid` package by Arkhangelsky et al. implements **Synthetic Difference-in-Differences**.
```r
library(synthdid)
# Prepare data as matrices
# Y: units × time matrix of outcomes
Y <- panel_data %>%
pivot_wider(id_cols = unit, names_from = time, values_from = outcome) %>%
column_to_rownames("unit") %>%
as.matrix()
# W: units × time matrix of treatment indicators (1 if treated)
W <- panel_data %>%
pivot_wider(id_cols = unit, names_from = time, values_from = treated) %>%
column_to_rownames("unit") %>%
as.matrix()
# Synthetic DiD estimate
tau_sdid <- synthdid_estimate(Y, W)
# Standard error (placebo-based)
se_sdid <- sqrt(vcov(tau_sdid, method = "placebo"))
# Compare to pure SC and pure DiD
tau_sc <- sc_estimate(Y, W)
tau_did <- did_estimate(Y, W)
# Visualize
synthdid_plot(tau_sdid)
```
## Worked Example
```{r}
#| label: worked-example
#| fig-cap: "Complete SCM analysis workflow"
# Create synthetic panel data for demonstration
set.seed(42)
n_units <- 15
n_periods <- 20
treatment_time <- 12
# Unit 1 is treated
panel <- expand.grid(
unit = 1:n_units,
time = 1:n_periods
) %>%
mutate(
treated_unit = (unit == 1),
post = (time >= treatment_time),
treatment = treated_unit & post,
# Pre-treatment characteristics
x1 = rnorm(n(), mean = unit/5, sd = 0.3),
x2 = rnorm(n(), mean = time/10, sd = 0.2),
# Outcome with treatment effect for unit 1
unit_fe = unit * 0.5,
time_fe = 0.15 * time,
tau = ifelse(treatment, -2.5 - 0.1*(time - treatment_time), 0),
y = unit_fe + time_fe + 0.3*x1 + 0.2*x2 + tau + rnorm(n(), 0, 0.4)
)
# --- Manual SCM Implementation ---
# Pre-treatment data
pre_data <- panel %>% filter(time < treatment_time)
# Treated unit pre-treatment outcomes
y_treated_pre <- pre_data %>%
filter(unit == 1) %>%
arrange(time) %>%
pull(y)
# Donor pre-treatment outcomes (matrix: time × donors)
Y_donors_pre <- pre_data %>%
filter(unit != 1) %>%
pivot_wider(id_cols = time, names_from = unit, values_from = y) %>%
arrange(time) %>%
select(-time) %>%
as.matrix()
# Find weights via constrained least squares (simplified)
# In practice, use quadprog or augsynth
# Here: OLS projection then normalize positive weights
weights_raw <- coef(lm(y_treated_pre ~ Y_donors_pre - 1))
weights_raw[is.na(weights_raw) | weights_raw < 0] <- 0
weights <- weights_raw / sum(weights_raw)
# Construct synthetic control for all periods
synthetic <- panel %>%
filter(unit != 1) %>%
left_join(
data.frame(unit = 2:n_units, weight = weights),
by = "unit"
) %>%
group_by(time) %>%
summarize(y_synth = sum(y * weight, na.rm = TRUE), .groups = "drop")
# Get treated outcomes
treated_outcomes <- panel %>%
filter(unit == 1) %>%
select(time, y_treated = y)
# Calculate gaps
results <- left_join(treated_outcomes, synthetic, by = "time") %>%
mutate(gap = y_treated - y_synth)
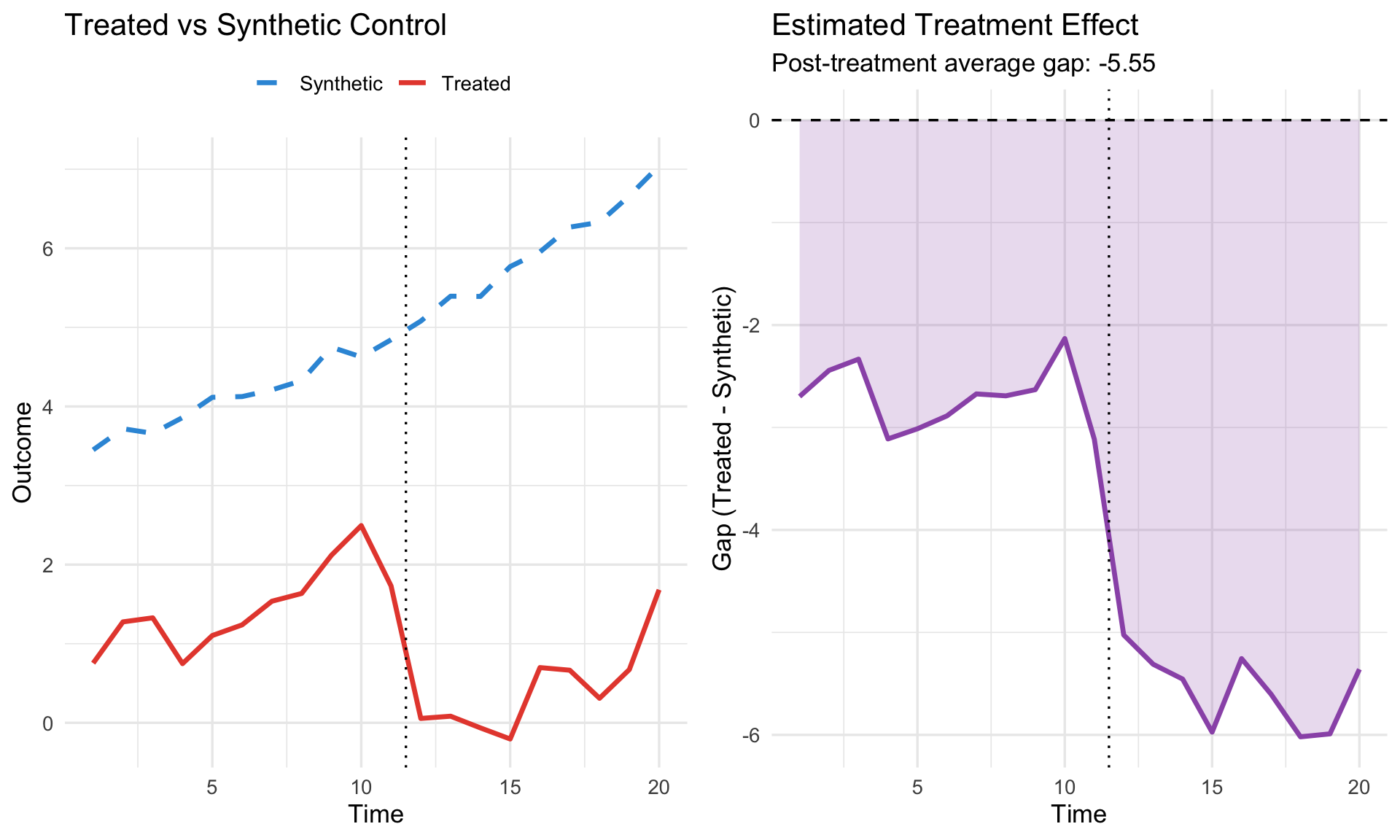
# --- Visualization ---
p1 <- ggplot(results, aes(x = time)) +
geom_line(aes(y = y_treated, color = "Treated"), linewidth = 1.2) +
geom_line(aes(y = y_synth, color = "Synthetic"), linewidth = 1.2, linetype = "dashed") +
geom_vline(xintercept = treatment_time - 0.5, linetype = "dotted") +
scale_color_manual(values = c("Treated" = "#e74c3c", "Synthetic" = "#3498db")) +
labs(title = "Treated vs Synthetic Control",
x = "Time", y = "Outcome", color = "") +
theme(legend.position = "top")
p2 <- ggplot(results, aes(x = time, y = gap)) +
geom_line(linewidth = 1.2, color = "#9b59b6") +
geom_ribbon(aes(ymin = pmin(0, gap), ymax = pmax(0, gap)),
fill = "#9b59b6", alpha = 0.2) +
geom_hline(yintercept = 0, linetype = "dashed") +
geom_vline(xintercept = treatment_time - 0.5, linetype = "dotted") +
labs(title = "Estimated Treatment Effect",
subtitle = paste0("Post-treatment average gap: ",
round(mean(results$gap[results$time >= treatment_time]), 2)),
x = "Time", y = "Gap (Treated - Synthetic)")
gridExtra::grid.arrange(p1, p2, ncol = 2)
```
# Diagnostics and Validation {.unnumbered}
## 1. Pre-Treatment Fit
**The most critical diagnostic**: How well does the synthetic control match the treated unit before treatment?
```{r}
#| label: pretreatment-fit
# Pre-treatment fit statistics
pre_fit <- results %>%
filter(time < treatment_time) %>%
summarize(
RMSPE = sqrt(mean(gap^2)),
MAE = mean(abs(gap)),
Max_Gap = max(abs(gap)),
Mean_Gap = mean(gap)
)
cat("Pre-Treatment Fit Diagnostics:\n")
cat(" RMSPE:", round(pre_fit$RMSPE, 3), "\n")
cat(" MAE:", round(pre_fit$MAE, 3), "\n")
cat(" Max Gap:", round(pre_fit$Max_Gap, 3), "\n")
cat(" Mean Gap:", round(pre_fit$Mean_Gap, 4), "(should be ~0)\n")
```
::: {.callout-warning}
## Poor Pre-Treatment Fit
If pre-treatment RMSPE is large (relative to the outcome's scale), the synthetic control is unreliable. Consider:
1. Adding more pre-treatment periods as predictors
2. Using augmented SCM (`progfunc = "Ridge"`)
3. Acknowledging the limitation in your analysis
:::
## 2. Weight Diagnostics
Check if weights are **sparse** and **sensible**.
```{r}
#| label: weight-check
# Weight distribution
weight_df <- data.frame(
unit = 2:n_units,
weight = weights
) %>%
filter(weight > 0.001) %>%
arrange(desc(weight))
cat("Donor Weights (non-zero):\n")
print(weight_df)
cat("\nNumber of donors with positive weight:", sum(weights > 0.001), "of", n_units - 1, "\n")
```
**Good signs**:
- Sparse weights (few donors contribute)
- Top donors make intuitive sense
- No single donor dominates completely
**Bad signs**:
- Many donors with tiny weights (overfitting)
- Counterintuitive donors receive high weight
- One donor receives weight ≈ 1 (just using that unit as control)
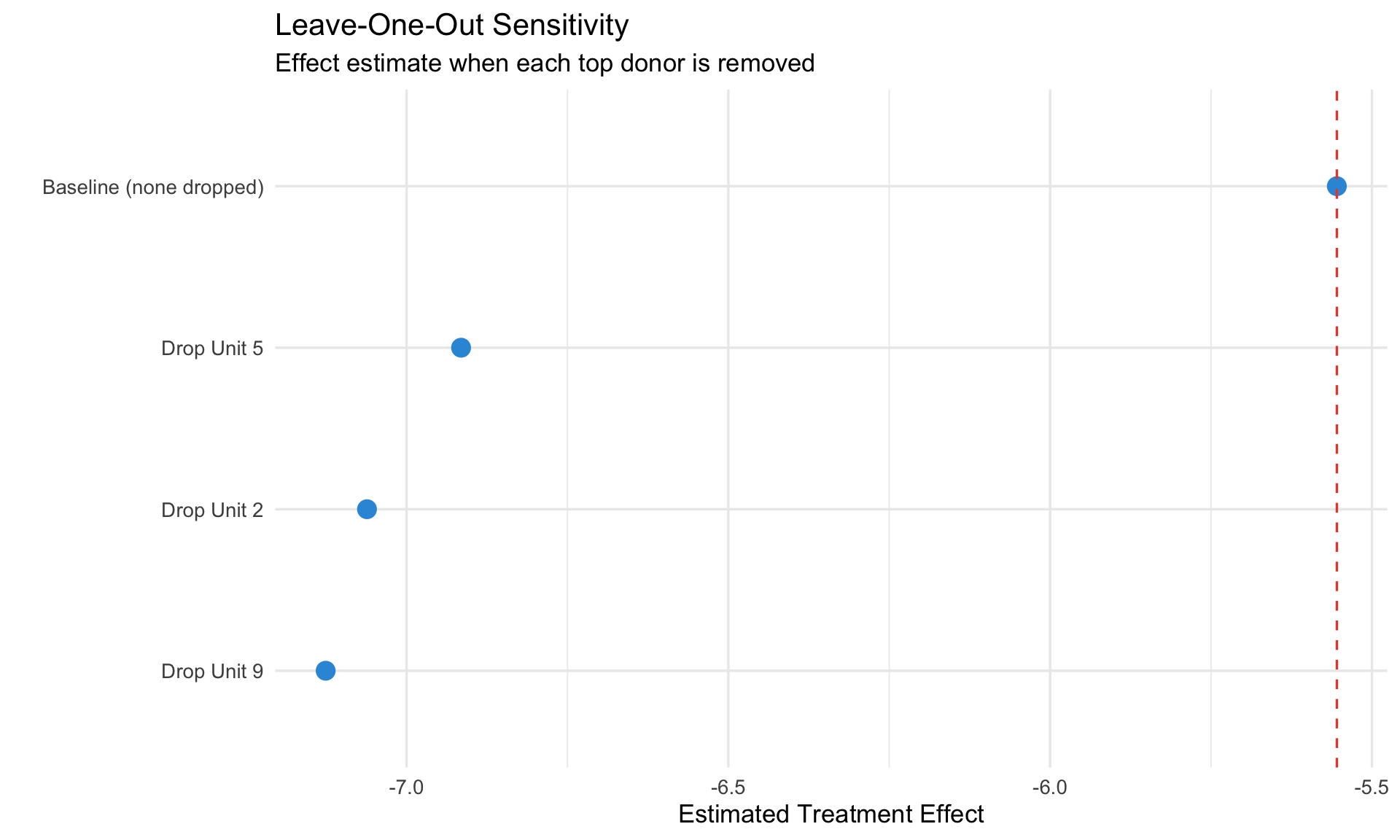
## 3. Leave-One-Out Sensitivity
Remove each important donor and re-estimate. If results are robust, the effect doesn't depend on any single donor.
```{r}
#| label: leave-one-out
#| fig-cap: "Leave-one-out sensitivity analysis"
# Identify top donors
top_donors <- weight_df$unit[1:min(3, nrow(weight_df))]
# Leave-one-out analysis
loo_results <- lapply(top_donors, function(drop_unit) {
# Re-estimate without this donor
Y_donors_loo <- pre_data %>%
filter(unit != 1, unit != drop_unit) %>%
pivot_wider(id_cols = time, names_from = unit, values_from = y) %>%
arrange(time) %>%
select(-time) %>%
as.matrix()
# New weights
w_loo <- coef(lm(y_treated_pre ~ Y_donors_loo - 1))
w_loo[is.na(w_loo) | w_loo < 0] <- 0
w_loo <- w_loo / sum(w_loo)
# New synthetic for post-treatment
remaining_units <- setdiff(2:n_units, drop_unit)
synth_loo <- panel %>%
filter(unit %in% remaining_units, time >= treatment_time) %>%
left_join(
data.frame(unit = remaining_units, weight = w_loo),
by = "unit"
) %>%
group_by(time) %>%
summarize(y_synth = sum(y * weight, na.rm = TRUE), .groups = "drop")
avg_gap <- mean(treated_outcomes$y_treated[treated_outcomes$time >= treatment_time] -
synth_loo$y_synth)
data.frame(dropped = paste("Drop Unit", drop_unit), effect = avg_gap)
}) %>% bind_rows()
# Add baseline
baseline <- mean(results$gap[results$time >= treatment_time])
loo_results <- bind_rows(
data.frame(dropped = "Baseline (none dropped)", effect = baseline),
loo_results
)
ggplot(loo_results, aes(x = reorder(dropped, effect), y = effect)) +
geom_point(size = 4, color = "#3498db") +
geom_hline(yintercept = baseline, linetype = "dashed", color = "#e74c3c") +
coord_flip() +
labs(title = "Leave-One-Out Sensitivity",
subtitle = "Effect estimate when each top donor is removed",
x = "", y = "Estimated Treatment Effect")
```
# SCM vs. DiD: Decision Framework {.unnumbered}
## When to Use Which?
```
How many treated units?
├── One treated unit
│ └── Use Synthetic Control
│ └── Enough donors with good pre-treatment fit?
│ ├── Yes → Classic SCM or Augmented SCM
│ └── No → Consider qualitative analysis
└── Multiple treated units
├── Same treatment timing?
│ ├── Yes → DiD or Synthetic DiD
│ └── No (staggered) → Staggered DiD methods (Module 5) or Synthetic DiD
└── Trust parallel trends?
├── Yes → Standard DiD
└── No → Synthetic DiD (reweights to match)
```
## Comparison Table
| Feature | DiD | SCM | Synthetic DiD |
|---------|-----|-----|---------------|
| **# Treated** | Many | One | One or many |
| **Parallel trends** | Required | Not required | Not required |
| **Pre-treatment fit** | Implicit | Explicit (visible) | Explicit |
| **Inference** | Standard | Permutation | Placebo/Bootstrap |
| **Transparency** | Moderate | High (weights) | High |
| **Covariates** | Easy | Via predictors | Via augmentation |
# Common Pitfalls {.unnumbered}
::: {.callout-warning}
## 1. Interpolation vs. Extrapolation
SCM works by **interpolation**—the synthetic unit must lie within the convex hull of donors. If the treated unit is extreme (outside the range of donors), SCM extrapolates and becomes unreliable.
**Check**: Are the treated unit's pre-treatment characteristics within the range of donors?
:::
::: {.callout-warning}
## 2. Overfitting Pre-Treatment Noise
With many pre-treatment periods and few donors, SCM can overfit noise rather than signal.
**Solutions**:
- Use augmented SCM (`progfunc = "Ridge"`)
- Check if weights are sensible
- Validate with leave-one-out
:::
::: {.callout-warning}
## 3. Small Donor Pool
With few donors (< 10), permutation inference has limited power. You may not be able to reject the null even with a real effect.
**Solution**: Be transparent about power limitations. Consider combining with qualitative evidence.
:::
::: {.callout-warning}
## 4. Spillovers / SUTVA Violation
If treatment affects donor units (e.g., trade diversion, policy spillovers), the synthetic control is contaminated.
**Solution**: Exclude donors likely affected by spillovers. Test sensitivity to donor pool composition.
:::
::: {.callout-warning}
## 5. Anticipation Effects
If units anticipate treatment and adjust behavior before the official treatment date, pre-treatment fit is compromised.
**Solution**: Use an earlier treatment date cutoff, or exclude periods with possible anticipation.
:::
# Summary {.unnumbered}
Key takeaways:
1. **SCM constructs counterfactuals** by weighting untreated units to match the treated unit's pre-treatment trajectory
2. **The optimization** minimizes pre-treatment prediction error subject to convex weights
3. **Inference via permutation**: Apply SCM to each donor as placebo; compare treated unit's gap to placebo distribution
4. **RMSPE ratio** normalizes by pre-treatment fit quality
5. **Synthetic DiD** extends SCM to multiple treated units without requiring parallel trends
6. **Key diagnostics**:
- Pre-treatment fit (RMSPE)
- Weight sparsity and sensibility
- Leave-one-out sensitivity
7. **Use SCM when**: one (or few) treated units, no natural control group, parallel trends questionable
8. **Software**: `augsynth` for augmented SCM, `synthdid` for Synthetic DiD
---
## Key References
**Foundational**:
- Abadie & Gardeazabal (2003). "The Economic Costs of Conflict." *AER*
- Abadie, Diamond & Hainmueller (2010). "Synthetic Control Methods for Comparative Case Studies." *JASA*
- Abadie, Diamond & Hainmueller (2015). "Comparative Politics and the Synthetic Control Method." *AJPS*
- Abadie (2021). "Using Synthetic Controls: Feasibility, Data Requirements, and Methodological Aspects." *JEL*
**Extensions**:
- Arkhangelsky et al. (2021). "Synthetic Difference-in-Differences." *AER*
- Ben-Michael, Feller & Rothstein (2021). "The Augmented Synthetic Control Method." *JASA*
- Doudchenko & Imbens (2016). "Balancing, Regression, Difference-in-Differences and Synthetic Control Methods." *NBER WP*
**Inference**:
- Firpo & Possebom (2018). "Synthetic Control Method: Inference, Sensitivity Analysis and Confidence Sets." *JCASP*
**R Packages**:
- `augsynth`: Ben-Michael et al. — Augmented synthetic control
- `synthdid`: Arkhangelsky et al. — Synthetic DiD
- `Synth`: Abadie et al. — Original SCM implementation
---
*Next: [Module 7: Bayesian Foundations](07_bayesian_foundations.qmd)*