---
title: "Regularization for Macro"
subtitle: "LASSO, Ridge, Elastic Net, and High-Dimensional Methods"
---
```{r}
#| label: setup
#| include: false
library(tidyverse)
library(glmnet)
set.seed(42)
```
# The High-Dimensional Challenge {.unnumbered}
Modern macroeconomics faces a **dimensionality problem**:
- Many potential predictors (hundreds of macro indicators)
- Short time series (T = 50-200 quarters)
- Traditional methods overfit: $K > T$ makes OLS infeasible
::: {.callout-note}
## The Core Trade-off
**Bias-variance trade-off**: More flexible models fit training data better but may generalize poorly. Regularization adds **bias** to reduce **variance**.
:::
## When Regularization Matters
| Setting | Problem | Solution |
|---------|---------|----------|
| **Large VARs** | K variables → K²p parameters | Minnesota prior (shrinkage) |
| **Forecasting** | Many predictors, short samples | LASSO/Ridge selection |
| **Factor models** | High-dimensional data | PCA, factor extraction |
| **Causal inference** | Many potential confounders | Double LASSO |
# Ridge Regression {.unnumbered}
## The Idea
Add an $\ell_2$ penalty to prevent coefficients from exploding:
$$
\hat{\beta}^{\text{ridge}} = \arg\min_\beta \left\{ \sum_{i=1}^n (y_i - x_i'\beta)^2 + \lambda \sum_{j=1}^p \beta_j^2 \right\}
$$
Equivalently:
$$
\hat{\beta}^{\text{ridge}} = (X'X + \lambda I)^{-1} X'y
$$
## Key Properties
| Property | Implication |
|----------|-------------|
| **Shrinks toward zero** | All coefficients are shrunk |
| **Never exactly zero** | Does NOT do variable selection |
| **Continuous** | Stable, smooth as function of $\lambda$ |
| **Handles multicollinearity** | $(X'X + \lambda I)$ always invertible |
```{r}
#| label: ridge-demo
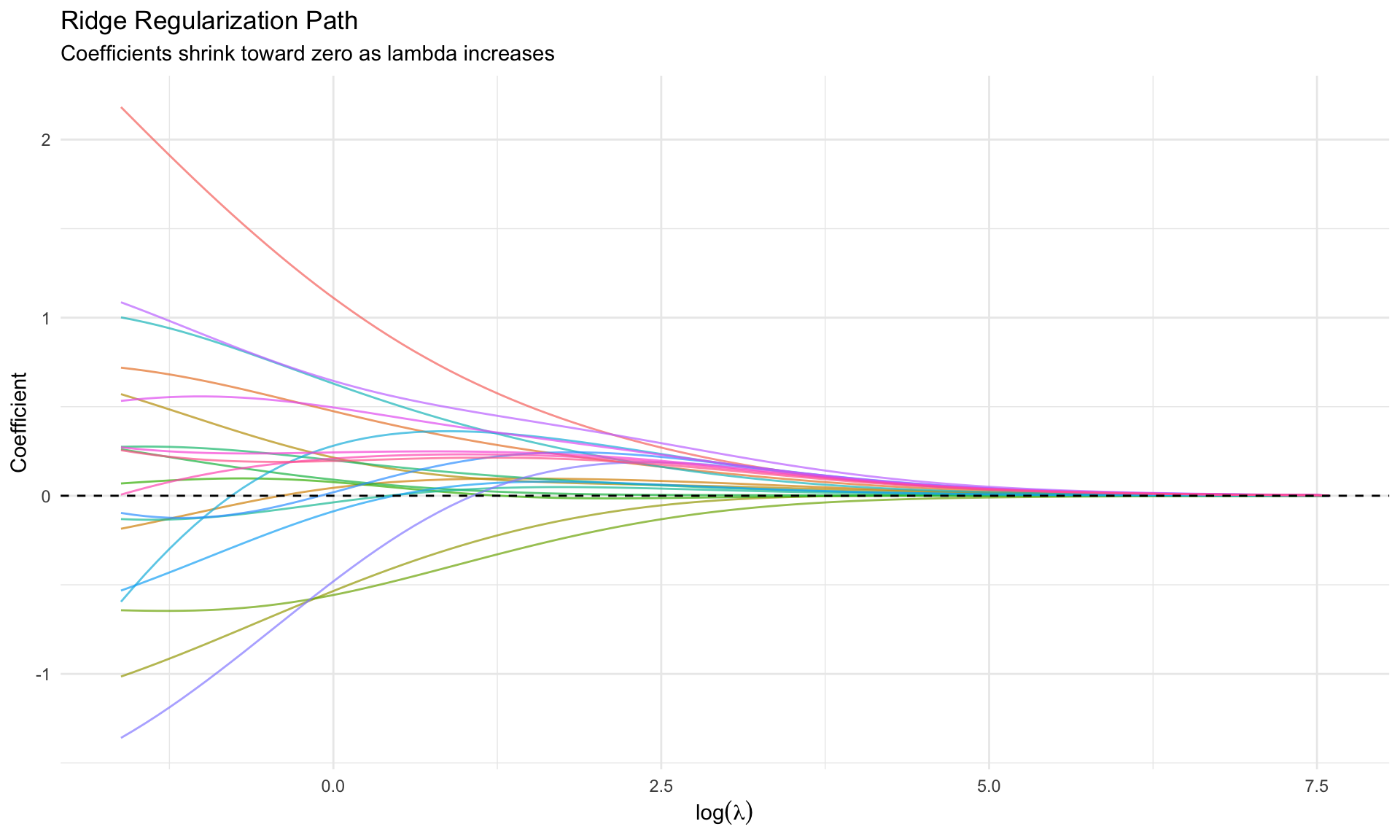
#| fig-cap: "Ridge regularization path"
# Simulate correlated predictors
n <- 100
p <- 20
X <- matrix(rnorm(n * p), n, p)
# Add correlation
for (j in 2:p) X[, j] <- 0.7 * X[, j-1] + 0.3 * X[, j]
# True coefficients: sparse
beta_true <- c(3, -2, 0, 0, 1.5, rep(0, p - 5))
y <- X %*% beta_true + rnorm(n, 0, 2)
# Ridge path
ridge_fit <- glmnet(X, y, alpha = 0) # alpha = 0 is Ridge
# Extract coefficients along lambda path
coef_matrix <- as.matrix(coef(ridge_fit)[-1, ]) # remove intercept
lambda_seq <- ridge_fit$lambda
ridge_df <- as_tibble(t(coef_matrix)) %>%
mutate(lambda = lambda_seq) %>%
pivot_longer(-lambda, names_to = "variable", values_to = "coefficient")
ggplot(ridge_df, aes(x = log(lambda), y = coefficient, color = variable)) +
geom_line(alpha = 0.7) +
geom_hline(yintercept = 0, linetype = "dashed") +
labs(title = "Ridge Regularization Path",
subtitle = "Coefficients shrink toward zero as lambda increases",
x = expression(log(lambda)), y = "Coefficient") +
theme_minimal() +
theme(legend.position = "none")
```
## Connection to Bayesian Priors
Ridge regression is equivalent to Bayesian regression with:
$$
\beta_j \sim N(0, \tau^2), \quad \tau^2 = \frac{\sigma^2}{\lambda}
$$
This is exactly the **Minnesota prior** intuition: shrink toward zero with tightness $\lambda$.
# LASSO {.unnumbered}
## The Idea
Replace $\ell_2$ penalty with $\ell_1$ (absolute value):
$$
\hat{\beta}^{\text{LASSO}} = \arg\min_\beta \left\{ \sum_{i=1}^n (y_i - x_i'\beta)^2 + \lambda \sum_{j=1}^p |\beta_j| \right\}
$$
## Key Properties
| Property | Implication |
|----------|-------------|
| **Shrinks to exactly zero** | Automatic variable selection |
| **Sparse solutions** | Most coefficients are zero |
| **Non-smooth** | Path has kinks (not differentiable) |
| **Selects one from correlated group** | Arbitrary among correlated predictors |
```{r}
#| label: lasso-demo
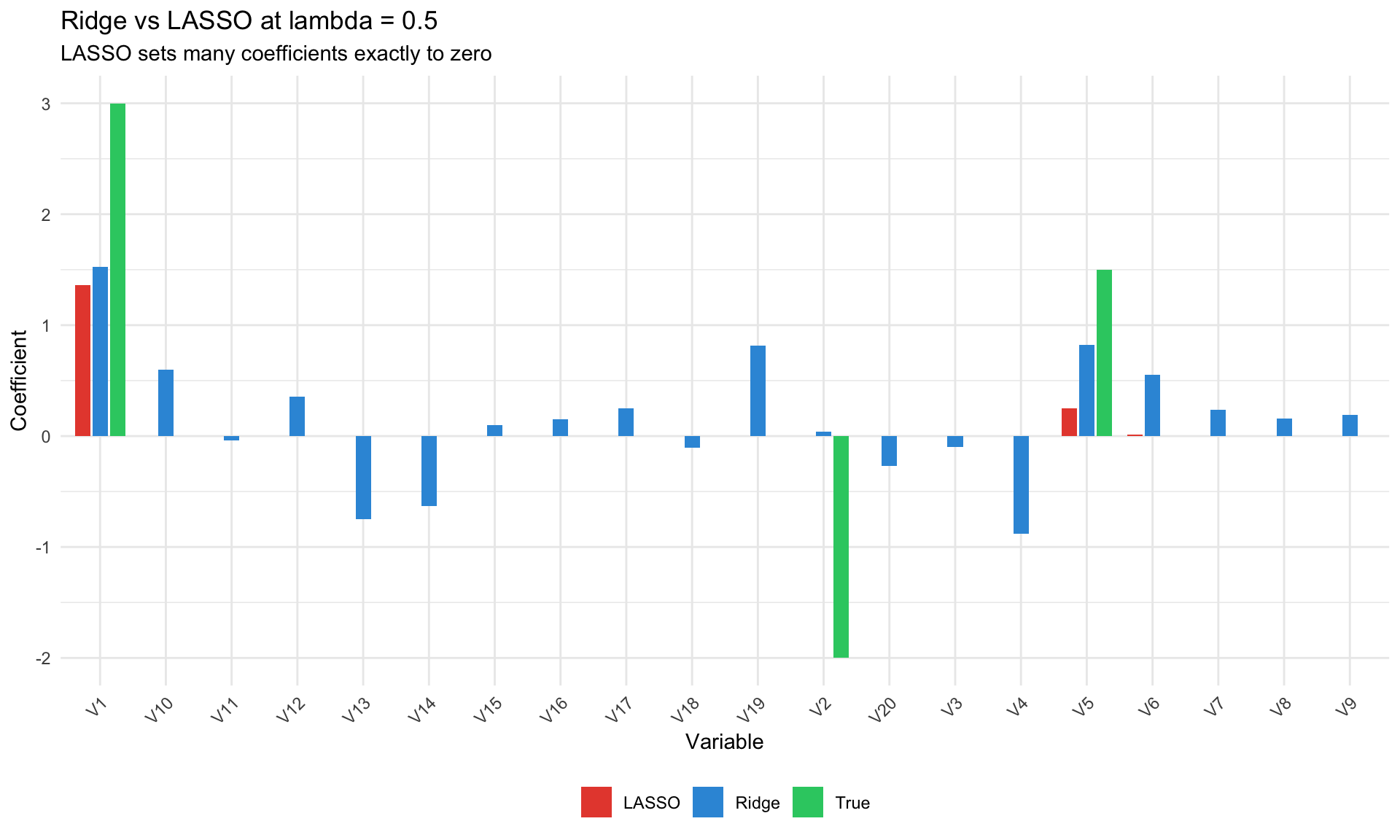
#| fig-cap: "LASSO vs Ridge: sparsity"
# LASSO path
lasso_fit <- glmnet(X, y, alpha = 1) # alpha = 1 is LASSO
coef_lasso <- as.matrix(coef(lasso_fit)[-1, ])
lambda_lasso <- lasso_fit$lambda
lasso_df <- as_tibble(t(coef_lasso)) %>%
mutate(lambda = lambda_lasso) %>%
pivot_longer(-lambda, names_to = "variable", values_to = "coefficient")
# Compare at specific lambda
lambda_compare <- 0.5
ridge_coef <- coef(ridge_fit, s = lambda_compare)[-1]
lasso_coef <- coef(lasso_fit, s = lambda_compare)[-1]
compare_df <- tibble(
variable = paste0("V", 1:p),
Ridge = as.vector(ridge_coef),
LASSO = as.vector(lasso_coef),
True = beta_true
) %>%
pivot_longer(c(Ridge, LASSO, True), names_to = "method", values_to = "coefficient")
ggplot(compare_df, aes(x = variable, y = coefficient, fill = method)) +
geom_bar(stat = "identity", position = position_dodge(width = 0.8), width = 0.7) +
scale_fill_manual(values = c("#e74c3c", "#3498db", "#2ecc71")) +
labs(title = paste0("Ridge vs LASSO at lambda = ", lambda_compare),
subtitle = "LASSO sets many coefficients exactly to zero",
x = "Variable", y = "Coefficient", fill = NULL) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1),
legend.position = "bottom")
```
```{r}
#| label: lasso-path
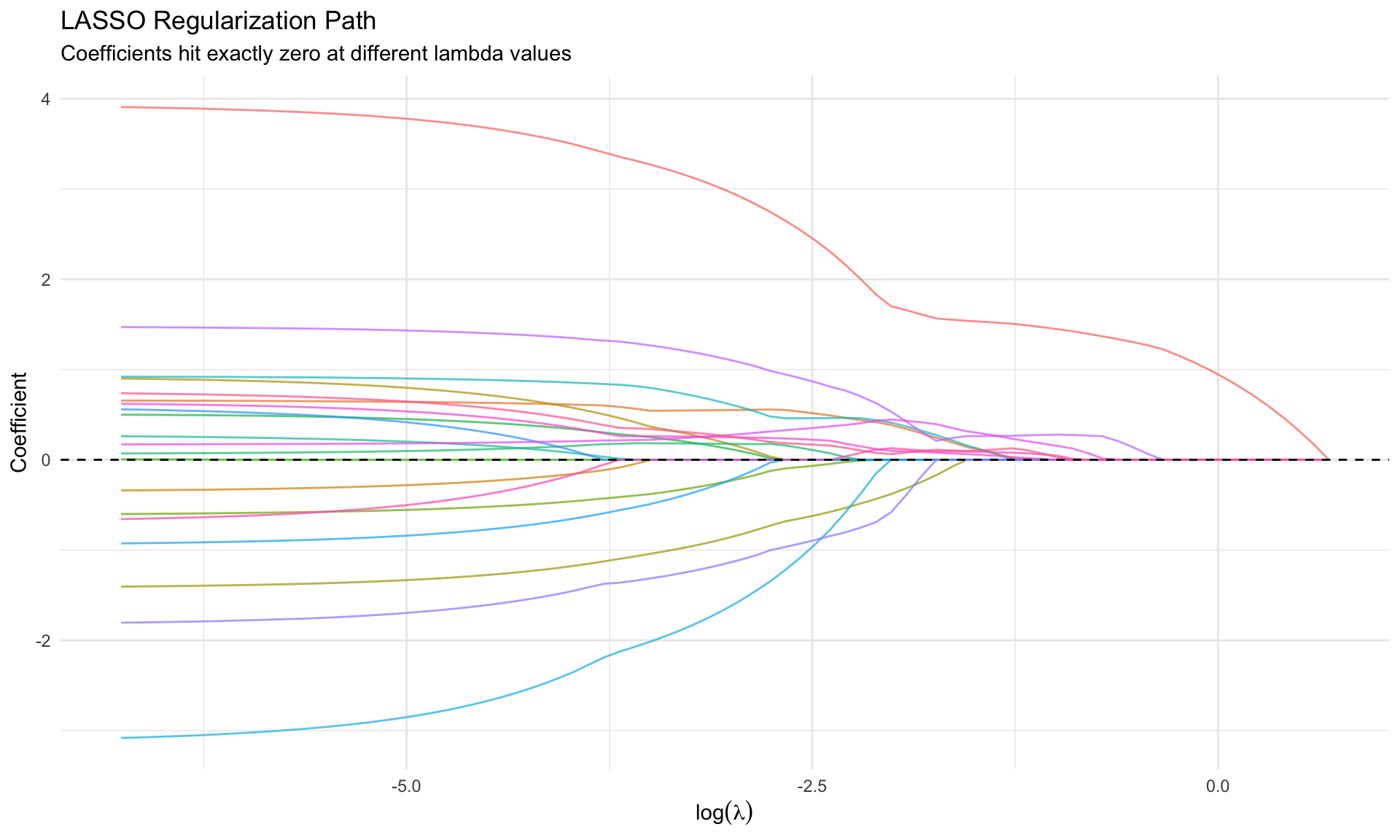
#| fig-cap: "LASSO path: coefficients drop to exactly zero"
ggplot(lasso_df, aes(x = log(lambda), y = coefficient, color = variable)) +
geom_line(alpha = 0.7) +
geom_hline(yintercept = 0, linetype = "dashed") +
labs(title = "LASSO Regularization Path",
subtitle = "Coefficients hit exactly zero at different lambda values",
x = expression(log(lambda)), y = "Coefficient") +
theme_minimal() +
theme(legend.position = "none")
```
# Elastic Net {.unnumbered}
## Combining Ridge and LASSO
$$
\hat{\beta}^{\text{EN}} = \arg\min_\beta \left\{ \sum_{i=1}^n (y_i - x_i'\beta)^2 + \lambda \left[ \alpha \sum_j |\beta_j| + (1-\alpha) \sum_j \beta_j^2 \right] \right\}
$$
| $\alpha$ | Method |
|----------|--------|
| 0 | Ridge |
| 1 | LASSO |
| 0.5 | Elastic Net (balanced) |
## When to Use Elastic Net
- **Correlated predictors**: LASSO arbitrarily selects one; Elastic Net keeps groups together
- **$p > n$**: LASSO selects at most $n$ variables; Elastic Net can select more
- **Default in practice**: Often use $\alpha = 0.5$ as a robust choice
```{r}
#| label: elastic-net
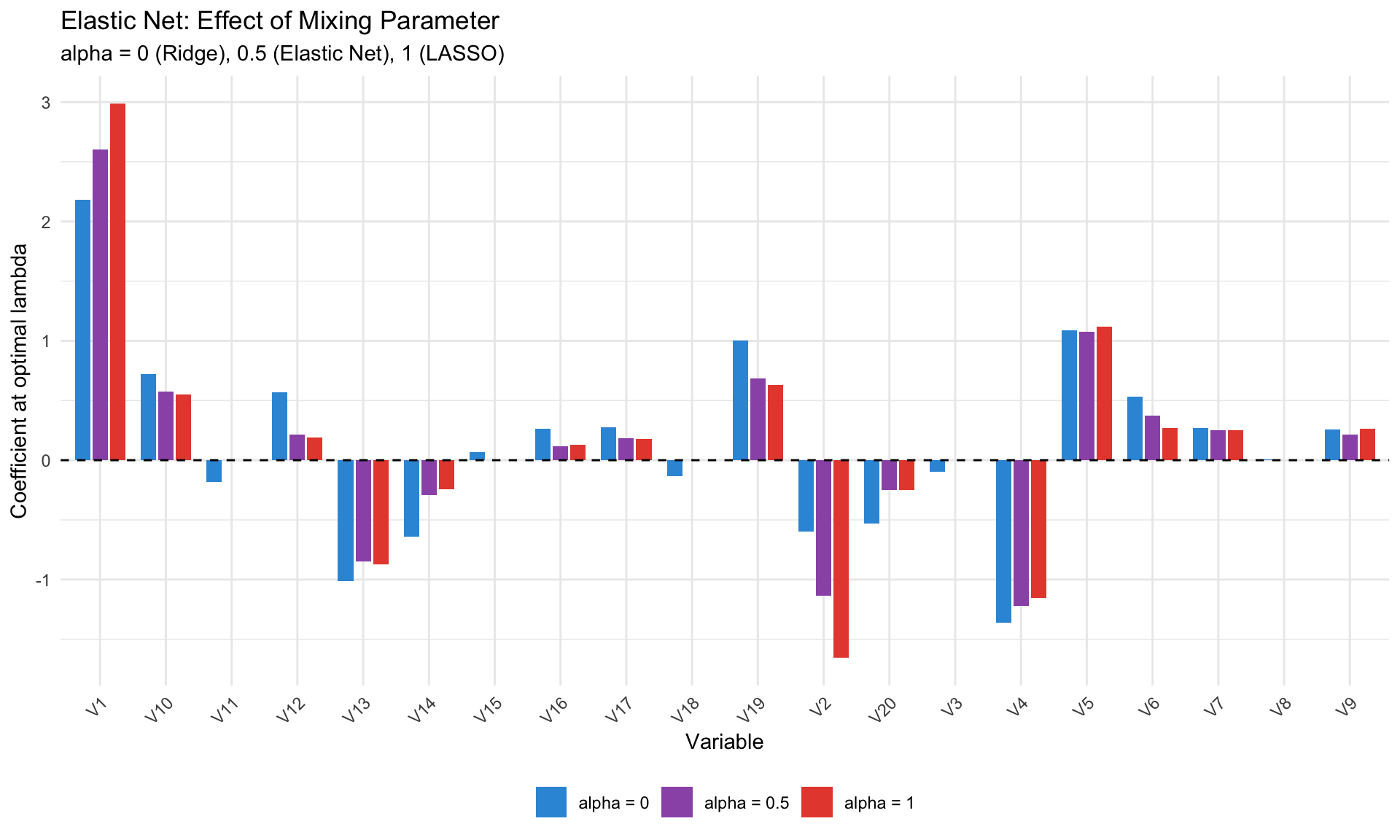
#| fig-cap: "Elastic Net for different alpha values"
alpha_values <- c(0, 0.5, 1)
en_fits <- map(alpha_values, ~glmnet(X, y, alpha = .x))
en_cv <- map(alpha_values, ~cv.glmnet(X, y, alpha = .x))
optimal_lambdas <- map_dbl(en_cv, ~.x$lambda.min)
en_coefs <- map2_dfr(en_fits, alpha_values, function(fit, a) {
lambda_opt <- optimal_lambdas[which(alpha_values == a)]
coef_vec <- coef(fit, s = lambda_opt)[-1]
tibble(
variable = paste0("V", 1:p),
coefficient = as.vector(coef_vec),
alpha = paste0("alpha = ", a)
)
})
ggplot(en_coefs, aes(x = variable, y = coefficient, fill = alpha)) +
geom_bar(stat = "identity", position = position_dodge(width = 0.8), width = 0.7) +
geom_hline(yintercept = 0, linetype = "dashed") +
labs(title = "Elastic Net: Effect of Mixing Parameter",
subtitle = "alpha = 0 (Ridge), 0.5 (Elastic Net), 1 (LASSO)",
x = "Variable", y = "Coefficient at optimal lambda", fill = NULL) +
scale_fill_manual(values = c("#3498db", "#9b59b6", "#e74c3c")) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1),
legend.position = "bottom")
```
# Choosing Lambda: Cross-Validation {.unnumbered}
## K-Fold Cross-Validation
1. Split data into K folds
2. For each $\lambda$:
- Train on K-1 folds, predict on held-out fold
- Repeat K times
- Average prediction error
3. Choose $\lambda$ that minimizes CV error
```{r}
#| label: cv-lambda
#| fig-cap: "Cross-validation for lambda selection"
# Cross-validation for LASSO
cv_lasso <- cv.glmnet(X, y, alpha = 1, nfolds = 10)
cv_df <- tibble(
log_lambda = log(cv_lasso$lambda),
cvm = cv_lasso$cvm,
cvup = cv_lasso$cvup,
cvlo = cv_lasso$cvlo
)
ggplot(cv_df, aes(x = log_lambda, y = cvm)) +
geom_point(color = "#e74c3c") +
geom_errorbar(aes(ymin = cvlo, ymax = cvup), alpha = 0.3) +
geom_vline(xintercept = log(cv_lasso$lambda.min), linetype = "dashed", color = "#3498db") +
geom_vline(xintercept = log(cv_lasso$lambda.1se), linetype = "dotted", color = "#2ecc71") +
annotate("text", x = log(cv_lasso$lambda.min) + 0.5, y = max(cv_df$cvm) * 0.9,
label = "lambda.min", color = "#3498db") +
annotate("text", x = log(cv_lasso$lambda.1se) + 0.5, y = max(cv_df$cvm) * 0.8,
label = "lambda.1se", color = "#2ecc71") +
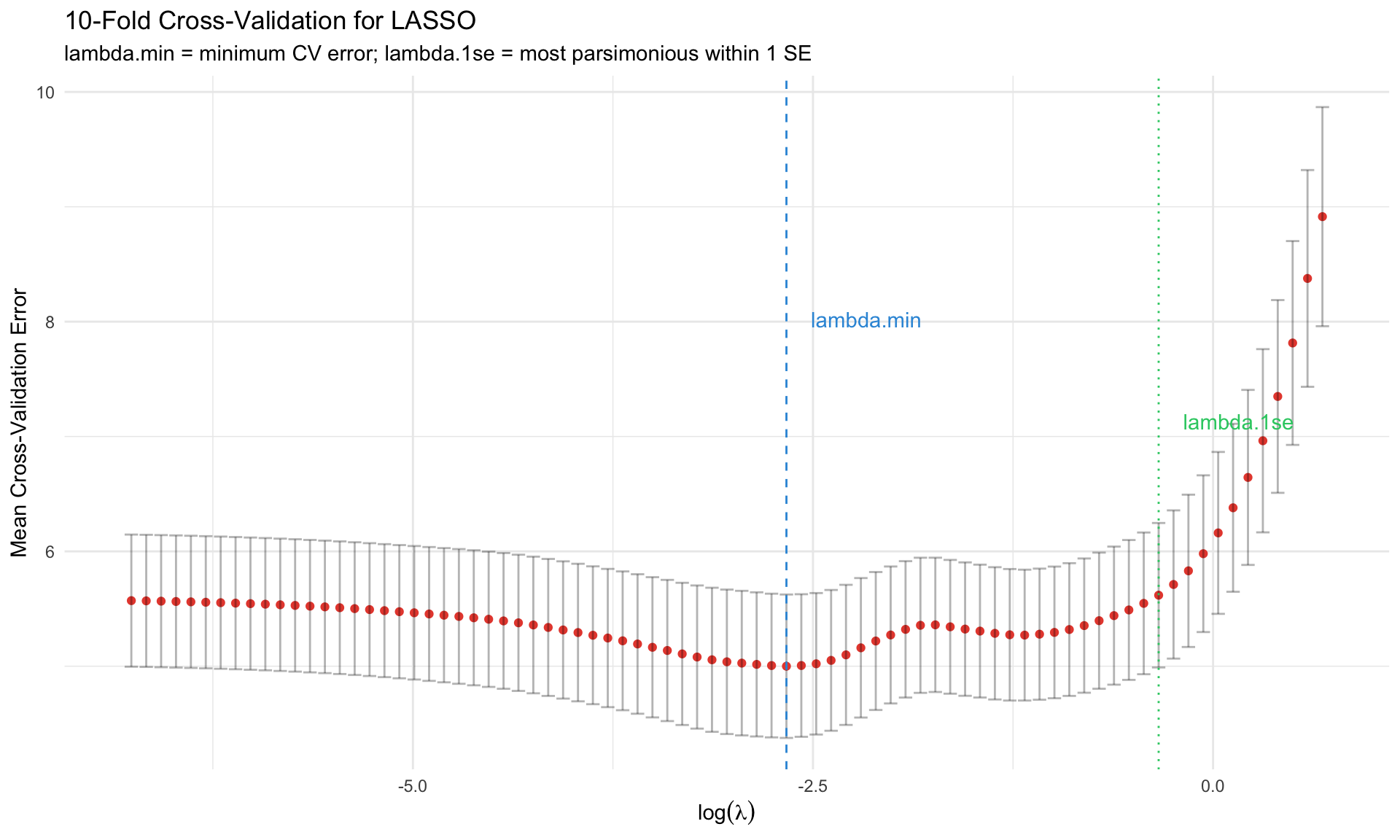
labs(title = "10-Fold Cross-Validation for LASSO",
subtitle = "lambda.min = minimum CV error; lambda.1se = most parsimonious within 1 SE",
x = expression(log(lambda)), y = "Mean Cross-Validation Error") +
theme_minimal()
```
## lambda.min vs lambda.1se
| Choice | Description | When to Use |
|--------|-------------|-------------|
| `lambda.min` | Minimizes CV error | Maximum predictive power |
| `lambda.1se` | Largest $\lambda$ within 1 SE of min | More parsimonious, often preferred |
# Double/Debiased LASSO {.unnumbered}
## The Problem with Naive LASSO for Inference
LASSO gives biased estimates and invalid standard errors because:
1. **Regularization bias**: Shrinkage toward zero
2. **Model selection uncertainty**: Which variables are "truly" zero?
3. **No valid confidence intervals** in the classical sense
## The Solution: Double Selection
For causal effect of $D$ on $Y$ with high-dimensional controls $X$:
**Step 1**: LASSO of $Y$ on $X$ → select controls $\hat{S}_Y$
**Step 2**: LASSO of $D$ on $X$ → select controls $\hat{S}_D$
**Step 3**: OLS of $Y$ on $D$ and $\hat{S}_Y \cup \hat{S}_D$
This is the **Belloni, Chernozhukov, Hansen (2014)** approach.
```{r}
#| label: double-lasso
#| fig-cap: "Double LASSO for causal inference"
# Simulate causal setting
n <- 200
p <- 50 # many potential confounders
# Generate confounders
X <- matrix(rnorm(n * p), n, p)
# Treatment depends on some confounders
D <- 0.5 * X[, 1] + 0.3 * X[, 2] + 0.2 * X[, 5] + rnorm(n, 0, 1)
# Outcome depends on treatment and confounders
tau_true <- 2 # true causal effect
Y <- tau_true * D + 1.5 * X[, 1] + 0.8 * X[, 3] + 0.5 * X[, 5] + rnorm(n, 0, 1)
# Naive OLS (omitting confounders)
naive_ols <- lm(Y ~ D)
cat("Naive OLS (biased):", round(coef(naive_ols)[2], 3), "\n")
# Double LASSO
# Step 1: Y on X
lasso_Y <- cv.glmnet(X, Y, alpha = 1)
selected_Y <- which(coef(lasso_Y, s = "lambda.1se")[-1] != 0)
# Step 2: D on X
lasso_D <- cv.glmnet(X, D, alpha = 1)
selected_D <- which(coef(lasso_D, s = "lambda.1se")[-1] != 0)
# Step 3: Union and OLS
selected_union <- unique(c(selected_Y, selected_D))
X_selected <- X[, selected_union, drop = FALSE]
double_lasso <- lm(Y ~ D + X_selected)
cat("Double LASSO:", round(coef(double_lasso)[2], 3), "\n")
cat("True effect:", tau_true, "\n")
cat("Selected variables:", length(selected_union), "\n")
# Compare estimates
comparison_df <- tibble(
Method = c("Naive OLS", "Double LASSO", "Oracle OLS"),
Estimate = c(
coef(naive_ols)[2],
coef(double_lasso)[2],
coef(lm(Y ~ D + X[, c(1, 3, 5)]))[2]
),
SE = c(
summary(naive_ols)$coefficients[2, 2],
summary(double_lasso)$coefficients[2, 2],
summary(lm(Y ~ D + X[, c(1, 3, 5)]))$coefficients[2, 2]
)
)
ggplot(comparison_df, aes(x = Method, y = Estimate)) +
geom_point(size = 4, color = "#3498db") +
geom_errorbar(aes(ymin = Estimate - 1.96 * SE, ymax = Estimate + 1.96 * SE),
width = 0.2, color = "#3498db") +
geom_hline(yintercept = tau_true, linetype = "dashed", color = "#e74c3c") +
annotate("text", x = 3.3, y = tau_true + 0.1, label = "True effect", color = "#e74c3c") +
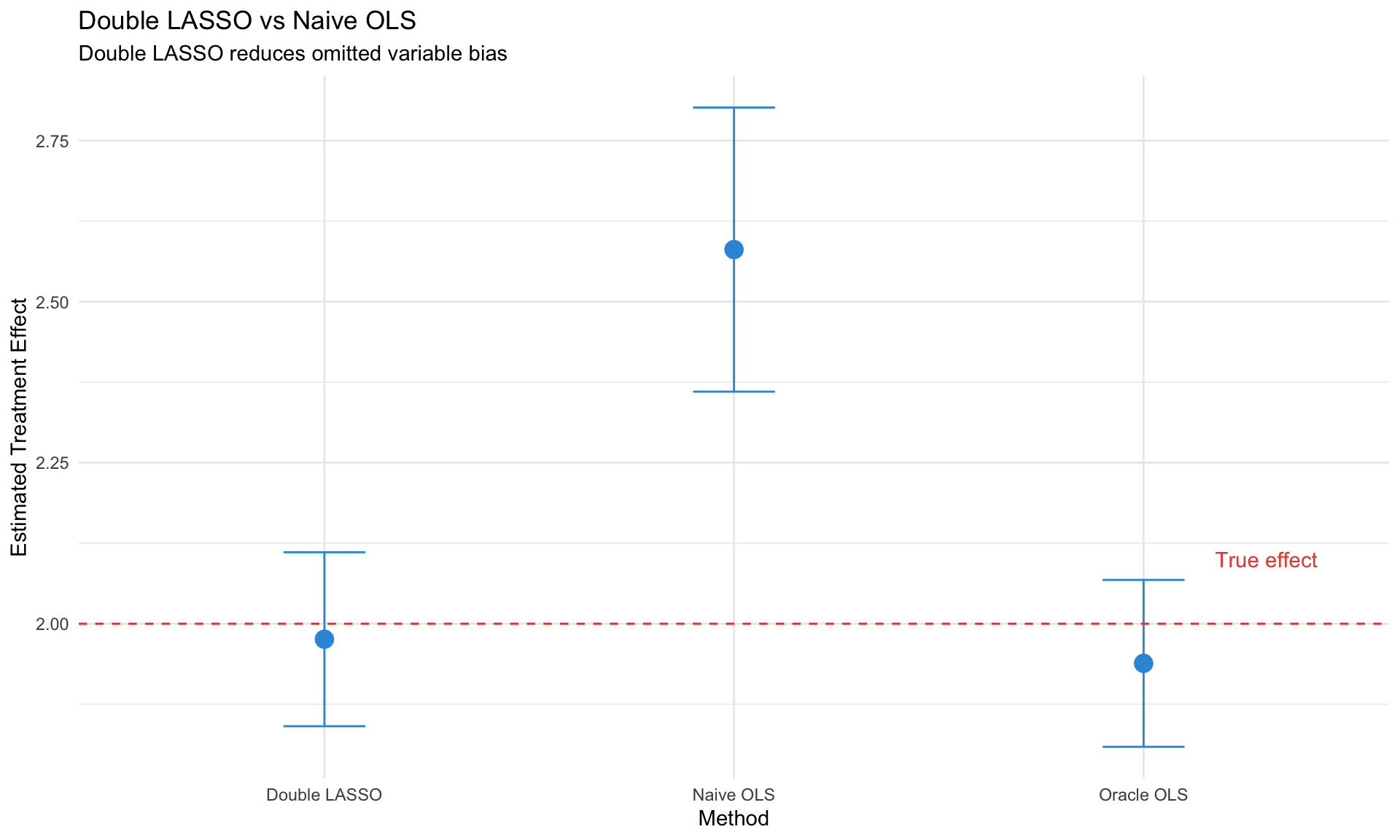
labs(title = "Double LASSO vs Naive OLS",
subtitle = "Double LASSO reduces omitted variable bias",
y = "Estimated Treatment Effect") +
theme_minimal()
```
## The `hdm` Package in R
```r
library(hdm)
# Double selection
ds_result <- rlassoEffect(x = X, y = Y, d = D, method = "double selection")
summary(ds_result)
# Partialling out
po_result <- rlassoEffect(x = X, y = Y, d = D, method = "partialling out")
summary(po_result)
```
# Applications in Macro {.unnumbered}
## Large Bayesian VARs (Banbura, Giannone, Reichlin 2010)
Problem: K-variable VAR(p) has $K^2 p$ parameters.
Solution: **Tighter Minnesota prior as dimension grows**
$$
\lambda_1^{\text{large}} = \lambda_1^{\text{small}} \times \frac{K_{\text{small}}}{K_{\text{large}}}
$$
This is implicit Ridge regularization.
## Factor-Augmented VAR (FAVAR)
Extract factors from large dataset, include in VAR:
$$
\begin{pmatrix} F_t \\ Y_t \end{pmatrix} = A \begin{pmatrix} F_{t-1} \\ Y_{t-1} \end{pmatrix} + u_t
$$
where $F_t$ are principal components of hundreds of macro series.
## Sparse Granger Causality
Use LASSO to identify which variables Granger-cause others in high-dimensional settings.
```{r}
#| label: sparse-granger
#| fig-cap: "Sparse VAR: identifying causal links"
# Simulate sparse VAR(1)
K <- 10 # variables
T_sim <- 200
# True VAR coefficient matrix (sparse)
A_true <- matrix(0, K, K)
A_true[1, 1] <- 0.7
A_true[2, 2] <- 0.6
A_true[3, 1] <- 0.3 # 1 → 3
A_true[4, 2] <- 0.25 # 2 → 4
A_true[5, 5] <- 0.8
# Simulate
Y_var <- matrix(0, T_sim, K)
for (t in 2:T_sim) {
Y_var[t, ] <- A_true %*% Y_var[t-1, ] + rnorm(K, 0, 0.5)
}
# Estimate with LASSO (equation by equation)
lasso_A <- matrix(0, K, K)
for (k in 1:K) {
y_k <- Y_var[-1, k]
X_lag <- Y_var[-T_sim, ]
cv_fit <- cv.glmnet(X_lag, y_k, alpha = 1)
lasso_A[k, ] <- coef(cv_fit, s = "lambda.1se")[-1]
}
# Compare
comparison_var <- tibble(
i = rep(1:K, each = K),
j = rep(1:K, K),
True = as.vector(t(A_true)),
LASSO = as.vector(t(lasso_A))
) %>%
filter(True != 0 | abs(LASSO) > 0.05)
ggplot(comparison_var, aes(x = True, y = LASSO)) +
geom_point(size = 3, alpha = 0.7) +
geom_abline(slope = 1, intercept = 0, linetype = "dashed", color = "#e74c3c") +
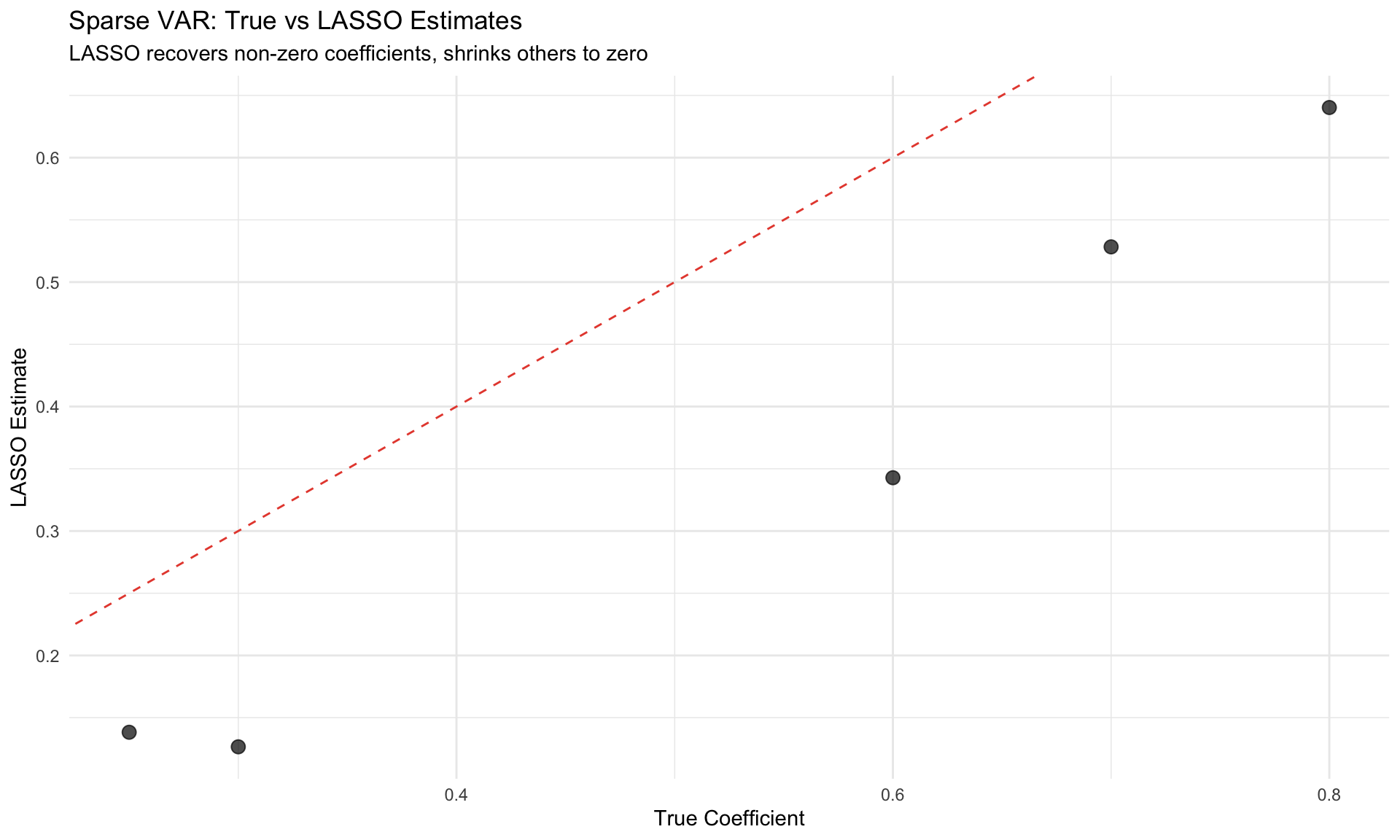
labs(title = "Sparse VAR: True vs LASSO Estimates",
subtitle = "LASSO recovers non-zero coefficients, shrinks others to zero",
x = "True Coefficient", y = "LASSO Estimate") +
theme_minimal()
```
# Practical Implementation {.unnumbered}
## The `glmnet` Workflow
```r
library(glmnet)
# Standardize predictors (glmnet does this internally)
X <- scale(X)
# Ridge (alpha = 0)
ridge_fit <- glmnet(X, y, alpha = 0)
cv_ridge <- cv.glmnet(X, y, alpha = 0)
coef(cv_ridge, s = "lambda.min")
# LASSO (alpha = 1)
lasso_fit <- glmnet(X, y, alpha = 1)
cv_lasso <- cv.glmnet(X, y, alpha = 1)
coef(cv_lasso, s = "lambda.1se")
# Elastic Net (alpha = 0.5)
en_fit <- glmnet(X, y, alpha = 0.5)
cv_en <- cv.glmnet(X, y, alpha = 0.5)
# Predictions
predict(cv_lasso, newx = X_new, s = "lambda.min")
```
## Choosing Alpha
Use nested cross-validation:
```r
# Grid search over alpha
alpha_grid <- seq(0, 1, 0.1)
cv_errors <- sapply(alpha_grid, function(a) {
cv_fit <- cv.glmnet(X, y, alpha = a)
min(cv_fit$cvm)
})
best_alpha <- alpha_grid[which.min(cv_errors)]
```
## Time Series Considerations
Standard CV is problematic for time series (breaks temporal structure).
**Solutions**:
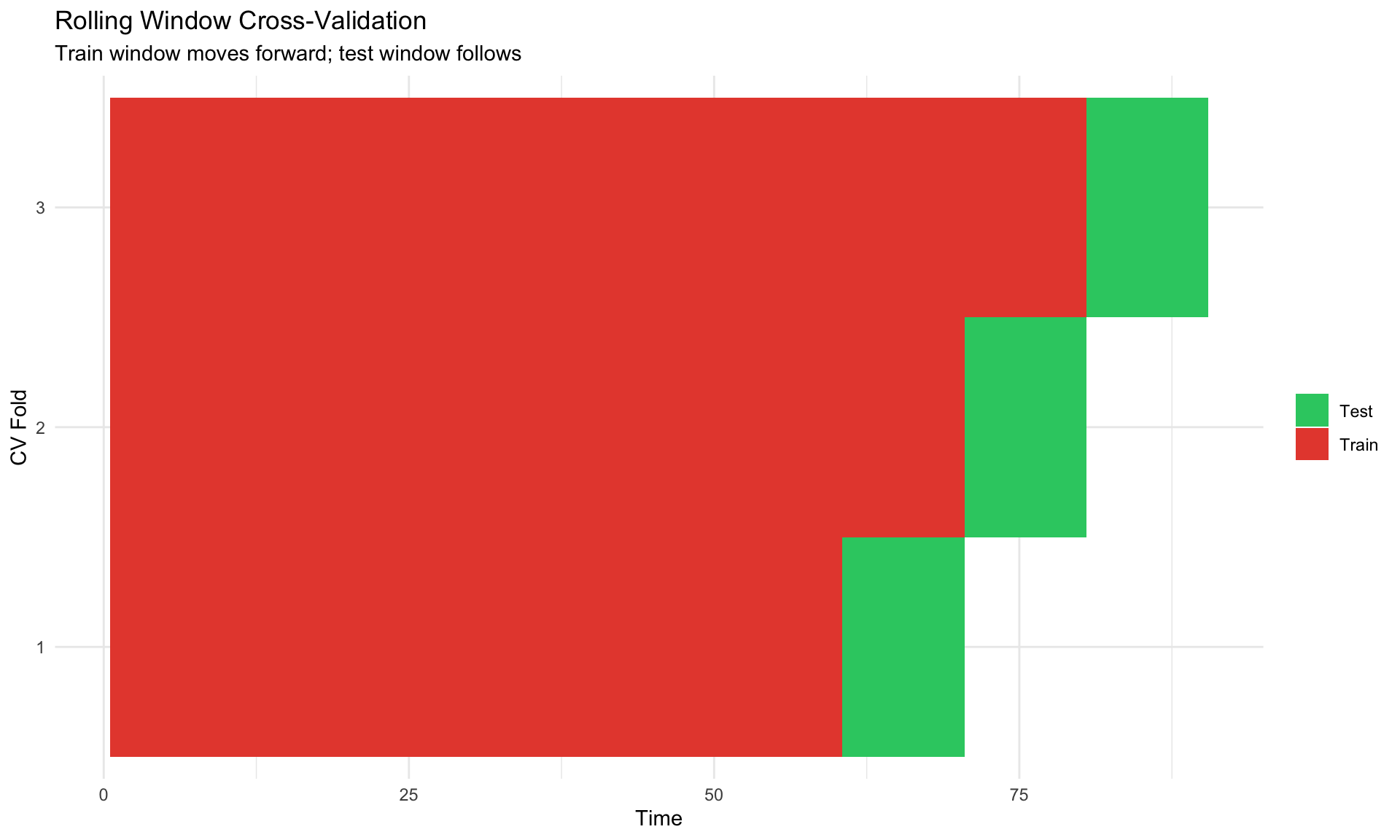
1. **Rolling window CV**: Train on $t=1,\ldots,s$, test on $s+1,\ldots,s+h$
2. **Expanding window CV**: Train on $t=1,\ldots,s$, test on $s+1$, then expand
3. **Block bootstrap**: Resample blocks to preserve autocorrelation
```{r}
#| label: ts-cv
#| fig-cap: "Time series cross-validation schemes"
# Illustrate rolling window CV
T_total <- 100
train_size <- 60
h <- 10 # horizon
cv_scheme <- tibble(
fold = rep(1:3, each = T_total),
t = rep(1:T_total, 3),
set = NA_character_
)
for (f in 1:3) {
train_end <- train_size + (f - 1) * h
test_end <- train_end + h
cv_scheme <- cv_scheme %>%
mutate(set = case_when(
fold == f & t <= train_end ~ "Train",
fold == f & t > train_end & t <= test_end ~ "Test",
TRUE ~ set
))
}
ggplot(cv_scheme %>% filter(!is.na(set)), aes(x = t, y = factor(fold), fill = set)) +
geom_tile() +
scale_fill_manual(values = c("#2ecc71", "#e74c3c")) +
labs(title = "Rolling Window Cross-Validation",
subtitle = "Train window moves forward; test window follows",
x = "Time", y = "CV Fold", fill = NULL) +
theme_minimal()
```
# Common Pitfalls {.unnumbered}
::: {.callout-warning}
## Watch Out For
1. **Post-selection inference**: Don't report OLS standard errors after LASSO selection
2. **Standardization**: Always standardize predictors for fair penalization
3. **Time series CV**: Don't use random splits for time series
4. **Interpretation**: LASSO coefficients are shrunken—not the "true" effects
5. **Correlated predictors**: LASSO picks arbitrarily; consider Elastic Net
6. **Oracle property**: LASSO is consistent under stringent conditions (irrepresentable condition)
:::
# Summary {.unnumbered}
| Method | Penalty | Selection | When to Use |
|--------|---------|-----------|-------------|
| **Ridge** | $\ell_2$ | No | Multicollinearity, all predictors matter |
| **LASSO** | $\ell_1$ | Yes | Sparse truth, variable selection |
| **Elastic Net** | Both | Yes | Correlated predictors, $p > n$ |
| **Double LASSO** | $\ell_1$ | Yes | Causal inference with many controls |
::: {.callout-tip}
## For Macro Applications
- **Forecasting**: LASSO/Ridge with rolling CV
- **Large VARs**: Minnesota prior (implicit Ridge)
- **Causal inference**: Double LASSO for high-dimensional controls
- **Model selection**: Elastic Net as robust default
:::
# Key References {.unnumbered}
## Regularization Methods
- **Tibshirani (1996)** "Regression Shrinkage and Selection via the Lasso" JRSS-B
- **Zou & Hastie (2005)** "Regularization and Variable Selection via the Elastic Net" JRSS-B
- **Hastie, Tibshirani & Wainwright (2015)** *Statistical Learning with Sparsity* — Comprehensive textbook
## High-Dimensional Inference
- **Belloni, Chernozhukov & Hansen (2014)** "Inference on Treatment Effects after Selection" RES — Double LASSO
- **Chernozhukov et al. (2018)** "Double/Debiased Machine Learning" Econometrics Journal
- **Athey & Imbens (2019)** "Machine Learning Methods That Economists Should Know About" Annual Review
## Macro Applications
- **Banbura, Giannone & Reichlin (2010)** "Large Bayesian VARs" JAE
- **Stock & Watson (2012)** "Generalized Shrinkage Methods for Forecasting" JAE
- **Giannone, Lenza & Primiceri (2021)** "Economic Predictions with Big Data" JAE
## Software
- **glmnet**: Friedman, Hastie & Tibshirani — R package for penalized regression
- **hdm**: Chernozhukov et al. — High-dimensional metrics in R